
circRNAs are circular noncoding RNAs formed by reverse splicing of pre-mRNAs. circRNAs were firstly found in viruses in the 1970s. However, due to the extensive use of the method for enrichment of poly (A) (no 5 'and 3' ends of circRNA) in the early RNA library preparation, and the calculation algorithm that RNA-seq reading requires linear alignment with genome, a large number of circRNA information was omitted, which led to the belief that circRNA is just a byproduct of miss splicing.
With the development of high-throughput sequencing technology and bioinformatics, thousands of circRNA have been found, and more and more basic researches related to circRNA have been done. A large number of studies have shown that circRNA is endogenetic, abundant, conservative and stable in mammalian cells, and often shows tissue or space-time specificity. It can participate in the regulation of cell growth and development, as well as the occurrence and development of diseases through a variety of mechanisms. Therefore, in recent years, circRNA has become popular in the field of non-coding RNA research.
Relationships between circRNAs and diseases
At present, the most studied is the relationship between circRNAs and tumors. Some circRNAs promote tumor formation, such as circPvt1 in squamous cell carcinomas of the head and neck, cirs-7 (CDr1as) in colorectal cancer, esophageal squamous cell carcinoma and hepatocellular carcinoma. Some circRNAs suppress tumors, such as circsMARCA5 and circ-SHPRH in glioblastoma. Some circRNAs may play different roles in different tissues or cells, such as circHiPK3, which is a proto-oncogene in rectal cancer, but suppresses cancer cells in bladder cancer.
In addition to cancers, circRNA has been found to be closely related to diabetes, cardiovascular disease, chronic inflammation and nervous system diseases. It is believed that with the development of biotechnology and more in-depth researches on circRNA, the formations and mechanisms of circRNAs can be identified. circRNAs can play important roles in disease prevention, diagnosis and treatment discovery.
The engineering and regulation of circRNA help disease studies and treatment discoveries
circRNA involves many complex functions, so how to study its functions? Similar to protein-coding genes, the most common methods are knockout, knockdown (RNA interference) or overexpression of the circRNA.
CRISPR/Cas9 mediated circRNA knockout, reveals the mechanism of its regulation on tumor formation
circRNA knockout refers to editing at the level of DNA to achieve the purpose of a complete knockout. The most commonly used method is to design two gRNAs at both ends of the circRNA exon to knockout the whole cyclized exon sequence. Although this strategy can knockout the circRNA, it will also affect the parent gene encoding the protein, and the study on its function is not ideal.
More and more researchers have given up using this strategy to knockout the circRNA because of its great influence on the parent gene. The ideal method is to knockout the loop forming elements (Alu) in the flanking intron of the exon, so as to destroy the circRNA loop forming without affecting the expression of the coding gene. However, it is difficult for many researchers to master the logic of designing the targeting strategy.
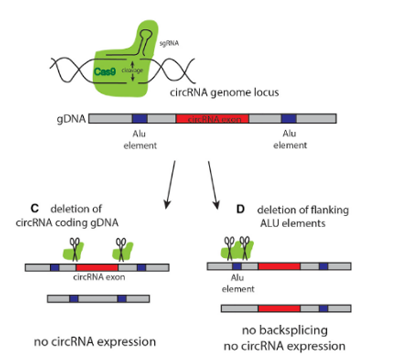
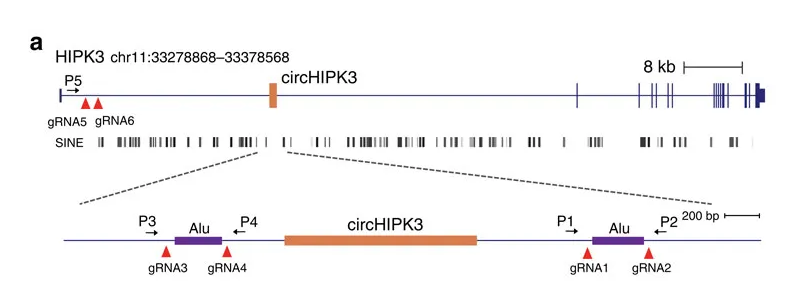
Taking circ-HIPK3 as an example, circ-HIPK3 is a kind of circRNA rich in human cells, which can combine with a variety of miRNAs as a regulator of cell growth and affect the formation of tumors. In order to verify how circ-HIPK3 forms into circle, it is necessary to find the loop forming elements in flanking intron. A pair of sgRNA is designed for the two Alu elements predicted at upstream and downstream respectively. The predicted loop forming elements are knockout by CRISPR/Cas9 system to detect whether the expression of circRNA changes. After PCR and RT-qPCR verification, it was found that the expression of circ-HIPK3 was significantly down-regulated after knockout of downstream loop forming elements, while the expression of circ-HIPK3 was not decreased but increased after knockout of upstream loop forming elements. It was speculated that there were too many loop forming elements in the upstream and the prediction was not accurate. In order to further verify the RNA circulation driven by other elements, the large fragment of intron in the upstream of the element was knockout by co-injection of gRNA3 or gRNA4 with gRNA5 or gRNA6. RT-qPCR results showed that the expression of circ-HIPK3 decreased, indicating that other loop forming elements of circ-HIPK3 exist.
Ubigene is experienced in designing strategy of knockout the loop forming elements in the flanking intron of circRNA exon, to achieve the purpose of knockout circRNA without affecting the expression of coding gene. Combined with CRISPR-UTM technology, the positive clones of circRNA knockout can be generated 10x faster than other common methods.
Knockdown specific circRNA and disclose its regulatory mechanism on cell proliferation and apoptosis
Among the methods to study the function of circRNA, the most classical way to inhibit circRNA is to knockdown it by RNAi (shRNA or siRNA). In order to avoid affecting mRNAs of coding genes, the shRNA should be designed at the back splicing site (BSS). siRNA was used to interfere with circ-HIPK3, and whether the knockdown of circ-HIPK3 would affect cell proliferation or apoptosis was observed. First, three groups of experiments targeting the linear transcript of HIPK3 mRNA, circ-HIPK3 circular transcript and both two transcripts. The designed siRNA interfered with the corresponding transcripts was verified on HEK-293T cell line. Cell proliferation and apoptosis were detected by CCK-8 and EdU assays. The results showed that the knockdown circ-HIPK3 significantly inhibited cell proliferation.
Ubigene can design high-score shRNA and use lentivirus to transfer the RNA interference vector into the cells. Cells were screened according to the drug screening, and the stable cell lines with circRNA knockdown were obtained.
Overexpression of circRNA reveal its loop-forming mechanism
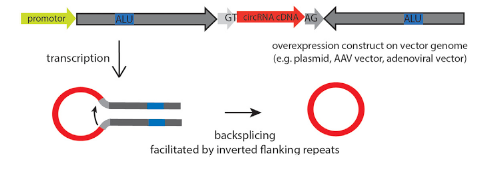
Overexpression of circRNA is not easy because of its low efficiency of loop forming, and it is easy to mismatch. By optimizing the binding sites of RBP, such as Alu elements and QKI, the circRNA can be formed accurately and efficiently. After overexpression, it is necessary to detect successful loop-forming and linear mRNA expression. In order to study the loop-forming efficiency of a new circRNA expression system, the mouse circrtn4 gene was selected to express in a variety of cell lines (including HeLa, N2a, HEK293). According to the RT-qPCR results in different cell lines, the efficiency of the new vector system pCircRNA-DMo-Rtn4 is much higher than that of the common vector system (pCircRNA-BE-Rtn4) in these cell lines.
CRISPR-U™ high-efficiency gene editing system
Ubigene focuses on genome editing, CRISPR-U™ is a gene-editing technology developed by Ubigene, which is more efficient than common CRISPR/Cas9 technologies in gene targeting. The services related to circRNA editing introduced in this paper can be provided by Ubigene, including knockout, interference, overexpression and circRNA expression testing.
References:
[1] Kristensen, L. S., Andersen, M. S., Stagsted, L. V., Ebbesen, K. K., Hansen, T. B., & Kjems, J. (2019). The biogenesis, biology and characterization of circular RNAs. Nature Reviews Genetics, 20(11), 675-691.
[2] Santer, L., Bär, C., & Thum, T. (2019). Circular RNAs, a novel class of functional RNA molecules with therapeutic perspective. Molecular Therapy.
[3] Zheng, Q., Bao, C., Guo, W., Li, S., Chen, J., Chen, B., ... & Liang, L. (2016). Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nature communications, 7(1), 1-13.



