CRISPR Screening Identifies OR7A10 as a Key Enhancer for CAR-NK Cell Therapy in Solid Tumors

Overview
Chimeric antigen receptor natural killer (CAR-NK) cell therapy holds considerable promise for the treatment of solid tumors. However, its clinical efficacy remains constrained by poor tumor infiltration, short persistence, and resistance within the tumor microenvironment (TME). A Yale University research team published a study in Nature that identified the GPCR family member OR7A10 as a core enhancer through in vivo CRISPR activation screening coupled with targeted ORF screening. CAR-NK cells engineered with OR7A10 cDNA, which bypasses the need for CRISPR editing and offers straightforward manufacturing, exhibit comprehensively enhanced functions, including increased proliferation, cytotoxicity, and metabolic fitness, alongside reduced exhaustion. These cells demonstrate potent efficacy across multiple solid tumour models (e.g., 100% complete response in an orthotopic breast cancer model), offering a highly potent and scalable off-the-shelf therapy for solid tumours.

Research Background
NK cells do not require MHC restriction or prior sensitisation and can target cancer cells with low mutational burden. CAR-NK therapy offers high safety and a low risk of GVHD, making it suitable for off-the-shelf manufacturing. Currently, there are over 1,200 clinical trials worldwide related to CAR-NK therapy, with more than 160 focused specifically on CAR-NK. However, this therapy faces core challenges in solid tumours, including insufficient tumour infiltration, poor proliferation, and poor persistence within the TME. Existing strategies, such as cytokine engineering or knockout of inhibitory genes, rely on CRISPR editing, which increases manufacturing complexity. Through dual in vivo screening, this study identified "enhancer" genes that can be directly integrated into the CAR construct, providing a simple and scalable solution for CAR-NK cell manufacturing.
Research Objectives
- To systematically identify genes (termed "super-enhancers" or "enhancers") that enhance the in vivo anti-tumour efficacy of CAR-NK cells via in vivo CRISPR activation screening and targeted ORF screening.
- To elucidate the molecular mechanisms by which OR7A10 enhances multiple functions of CAR-NK cells.
- To validate the in vivo efficacy of OR7A10-engineered CAR-NK cells across multiple solid tumour models.
- To evaluate the clinical translation potential of OR7A10-engineered CAR-NK cells, including manufacturing feasibility and safety.
- To explore the synergistic effects of OR7A10 with other activating signals, providing a theoretical foundation for optimising CAR-NK cell therapy.
Research Methods
Dual Screening Strategy
- In vivo CRISPRa screening: HER2-CAR-NK92 cells were transduced with a genome-wide CRISPR activation (CRISPRa) library based on the SAM system and adoptively transferred into HT29 colorectal tumor-bearing mice. Tumor-infiltrating cells were collected and compared with pre-infusion cells using next-generation sequencing (NGS) to analyze sgRNA abundance differences, enabling the identification of candidate genes that enhance CAR-NK cell activity.
-
In vivo barcoded ORF mini-library screening:
Primary human peripheral blood NK (PBNK) cells were first engineered to express HER2-CAR and subsequently transduced with the ORF library. These cells were infused into tumor-bearing mice, and tumor tissues were harvested at days 3 and 6 post-infusion. NGS was performed to quantify ORF and unique molecular identifier (UMI) barcode abundance, allowing prioritization of functional enhancer genes.
Functional Validation and Mechanistic Dissection
OR7A10 overexpression (OR7A10 (OE)) and stop-mutant (OR7A10 (STOP)) cell lines were generated. Cellular functions were assessed using flow cytometry, co-culture assays, Seahorse metabolic analysis, and TEM. CRISPR-Cas9 knockout of the GNAS gene, combined with small-molecule inhibitors, was employed to explore GPCR signalling pathways. Bulk mRNA-seq and single-cell transcriptomics were used to analyse gene expression profiles.
In Vivo Efficacy and Safety Validation
Antitumor efficacy was evaluated in HT29 colorectal cancer, MCF7 breast cancer, and SKOV3 ovarian cancer models through tumour volume measurements, bioluminescence imaging, and survival curves. Safety was assessed by measuring serum cytokine levels, performing histopathological analysis, and evaluating genomic integrity via WGS.
Research Workflow
Genome-wide CRISPRa screen → Targeted ORF screen identifying OR7A10 → In vitro functional validation (proliferation, cytotoxicity, etc.) → Mechanistic dissection (signalling pathways, gene expression) → In vivo efficacy and safety validation → Exploration of synergistic effects.
Key Findings
1. Dual in vivo screening identifies OR7A10 as a Core Enhancer
The CRISPRa screen identified 66 candidate genes, and the ORF mini-screen validated OR7A10 as the only significantly enriched gene, with its UMI showing strong enrichment in tumour samples.
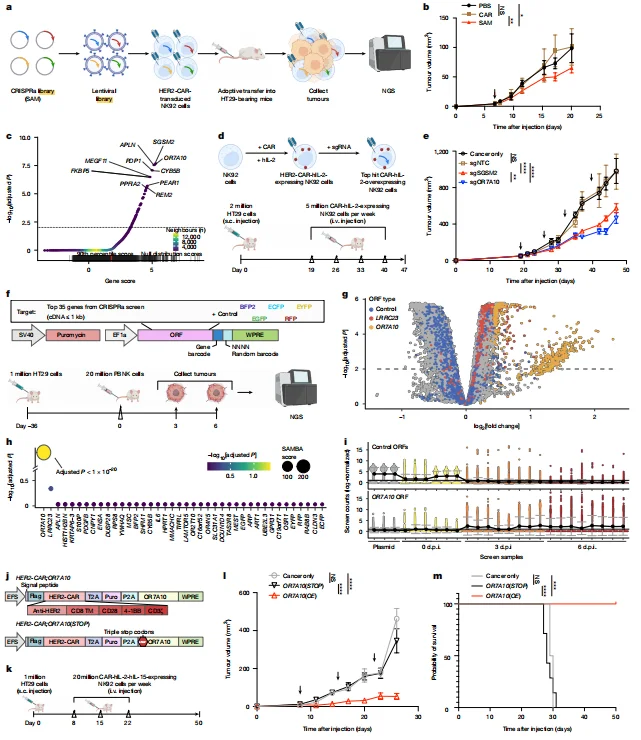
Figure 1. In vivo CRISPRa and barcoded ORF screens identify OR7A10 as an enhancer of CAR-NK cell anti-tumour efficacy.
2. OR7A10 enhances in vivo efficacy of CAR-NK cells
CAR-NK cells transfected with OR7A10 (OE) significantly suppressed tumour growth in HT29, MCF7 and SKOV3.In the MCF7 model, treatment achieved 100% complete response, accompanied by enhanced tumor infiltration and prolonged in vivo persistence of CAR-NK cells.
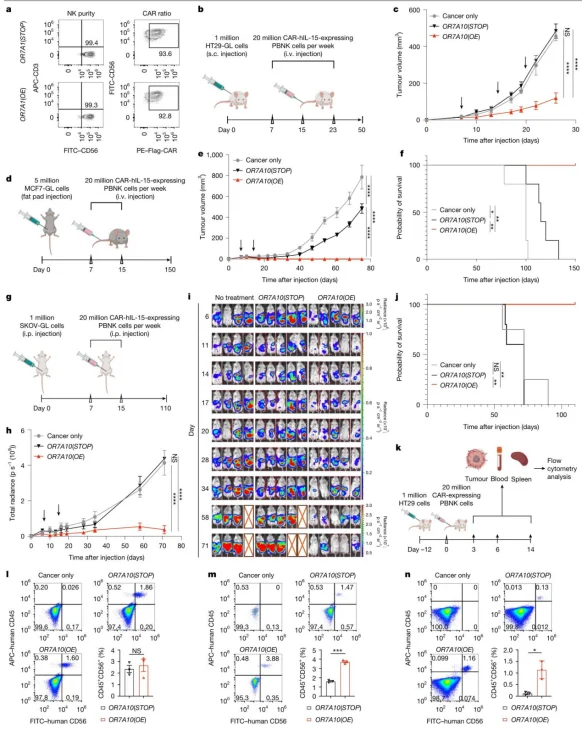
Figure 2. CAR-PBNK cells engineered with the OR7A10 open reading frame (ORF) exhibit robust in vivo anti-tumour activity and enhanced tumour infiltration across multiple tumour models.
3. Enhancement of multiple effector functions of primary CAR-NK cells
OR7A10 (OE)-transfected PBNK cells showed significantly enhanced cytotoxicity against various cancer cells, increased degranulation, cytokine (IFNγ, TNF) production, and death ligand (FasL, TRAIL) expression. Proliferation capacity was enhanced, while exhaustion markers (TIM-3, LAG-3, etc.) were reduced.
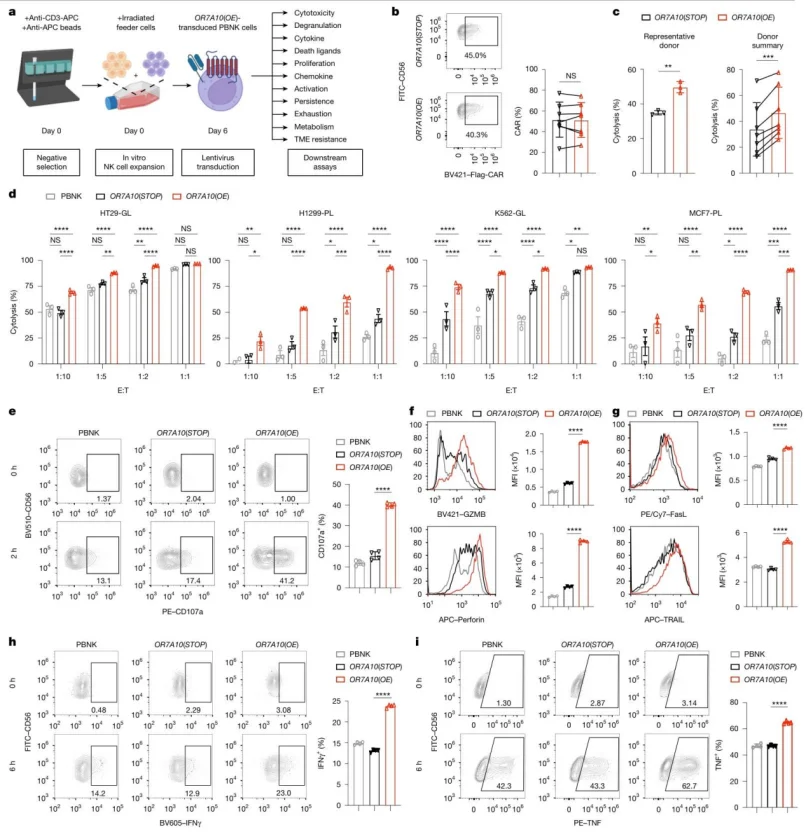
Figure 3. OR7A10 ORF engineering enhances the anti-tumour function of CAR-PBNK cells.
4. Improved metabolic fitness and resistance to the TME
OR7A10 (OE) CAR-NK cells displayed enhanced metabolic capacity, characterized by increased basal, spare, and maximal oxygen consumption rates, as well as elevated mitochondrial mass. Under immunosuppressive conditions such as acidic pH, TGFβ, and hypoxia, these cells maintained significantly higher cytotoxicity compared to control groups.
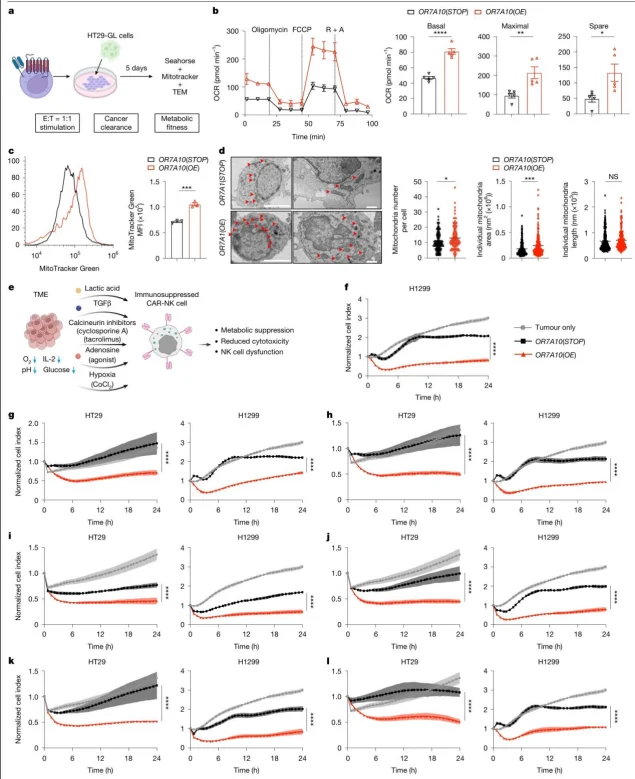
Figure 4. OR7A10 ORF engineering enhances the metabolic fitness of CAR-PBNK cells and their tolerance to TME-related immunosuppression.
5. Single-cell transcriptomics reveals regulatory features
OR7A10 overexpression exerted broad effects across all eight NK cell subsets, upregulating genes associated with inflammatory cytokines, cytotoxicity, and activating pathways such as NFAT and JNK.
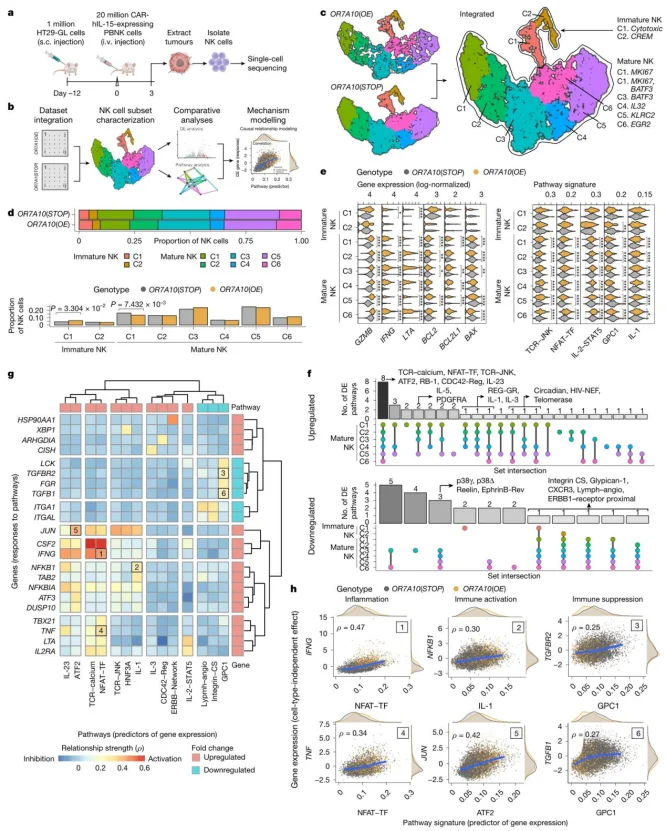
Figure 5. Single-cell transcriptomics (sCT) reveals unbiased in vivo gene expression changes driven by OR7A10 overexpression in tumour-infiltrating primary human CAR-NK cells.
Summary
This study identified the GPCR molecule OR7A10 through dual in vivo screening. Its overexpression comprehensively enhances the proliferation, cytotoxicity, metabolic fitness, tumour infiltration capacity, and TME resistance of CAR-NK cells while reducing exhaustion. Mechanistically, OR7A10 relies on GPCR-related signalling pathways to regulate gene expression profiles. OR7A10-engineered CAR-NK cells are straightforward to manufacture, demonstrate remarkable efficacy across multiple solid tumour models with a favourable safety profile and no risk of GVHD, and exhibit synergistic effects with NKp46 signalling. This research deepens the understanding of CAR-NK cell functional regulation and provides a highly potent, scalable, and safe CAR-NK cell therapy for solid tumours, offering significant clinical translation potential.
CRISPR-iScreen™ Libraries from Ubigene
CRISPR-iScreen™ is an innovative technology independently developed by Ubigene, designed to achieve efficient CRISPR screening. Currently, Ubigene offers over 40 CRISPR library Plasmids in stock, covering genome-wide knockout/Interference/activation libraries for humans, mice, green monkeys, pigs, and other species. Additionally, there are CRISPR knockout/Interference/activation sub-libraries targeting genes related to kinases, the cell cycle, membrane proteins, and metabolism, helping you easily identify targets of interest! Ubigene currently has over 150 CRISPR library virus stocks and over 600 CRISPR library Cell Pools, and also provides one-stop CRISPR (in vitro/in vivo) functional screening services to readily support your research needs.
Contact us to learn more>>>


