Gene Point Mutation Cell Line
Gene point mutation cell lines are generated by co-delivering, via nucleofection, ribonucleoprotein (RNP) complexes composed of gRNA and Cas9 protein together with a single-stranded oligonucleotide (ssODN) donor harboring the desired point mutation into cells. Subsequently, single-cell clones are isolated using the limiting dilution method. Following the induction of DNA double-strand breaks (DSBs) at the target locus by the RNP complex, cells utilize the ssODN donor as a repair template to perform homology-directed repair (HDR), thereby precisely incorporating the intended point mutation into the genomic target site. The target region is then amplified and validated by sequencing to identify and screen positive mutant clones.
Point Mutation Cell Line Service Detail
Service Type
Gene Point Mutation
Cell Line Type
Tumor Cells, Immortalized Cell Lines. Various cell types, including iPS cells
Project Price (USD)
From $4980
Turnaround
As fast as 6 weeks
Deliverables
1. ≥1 homozygous Single Clone(>106 cells/vial, 2 vials per clone) 2. Wild-Type Cells(>106 cells/vial, 2 vials) 3. Experimental Reports
Point mutation validation
PCR(Standard), Sanger sequencing(Standard)
Digestive System
Endocrine System
Respiratory System
Reproductive System
Circulatory System
Blood and lymphatic System
Brain and Nervous System
Urinary System
Skeleton, Articulus, Soft Tissue, Derma System
Stem Cell Lines
Ocular, Otolaryngologic and Oral System

Point Mutation (PM) Cells Construction Methods
4 solutions to meet different mutation needs!
Incubate sgRNA and Cas9 protein in vitro to form an RNP (ribonucleoprotein) complex, which is then co-transfected into cells along with a single-stranded oligonucleotide. After the RNP induces a double-strand break at the target site, the oligonucleotide can introduce the desired mutation into the genomic target through homologous recombination.
- Broad applicability
- High editing efficiency
- Short turnaround time
Features:
- Suitable when a gRNA sequence can be identified within approximately 10 bp of the mutation site.
- Applicable to target cells that can be transfected using electroporation or liposome-based transfection.
Applicable Types:
Gene editing is achieved by fusing an engineered reverse transcriptase with a Cas9 nickase and using a special prime editing guide RNA (pegRNA).
- Capable of correcting various complex mutations.
- However, it has low editing efficiency and requires complex vector design.
Features:
- Suitable for correcting complex mutations.
- Ideal for cases where an appropriate gRNA sequence cannot be found near the mutation site.
Applicable Types:
A base editing system is developed by integrating a base deaminase with the CRISPR/Cas system, enabling precise introduction of C/G-to-T/A and A/T-to-G/C point mutations without creating double-strand breaks, thereby achieving highly efficient and accurate gene editing.
- High efficiency, but applicable cases are limited.
Features:
- Suitable for specific single-base conversions (C⇌T; A⇌G) when the target site is within the active window.
Applicable Types:
The donor vector is constructed using a plasmid, with a resistance gene expression cassette inserted into an intron to enhance the efficiency of point mutation recombination through antibiotic screening.
- Suitable for scenarios where a suitable gRNA sequence cannot be identified near the mutation site; however, it is relatively expensive and has a longer experimental cycle.
Features:
- Ideal for cases where a suitable gRNA sequence cannot be identified near the mutation site.
Applicable Types:
Workflow and Validation
Strategy Design by Red Cotton System

RNP Complex
Cell Transfection

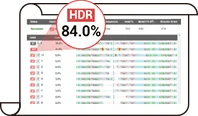
Pool Efficiency Validation
Single-cell Cloning
PCR Amplification
Sanger Sequencing Validation
QC & Cell Cryopreservaion

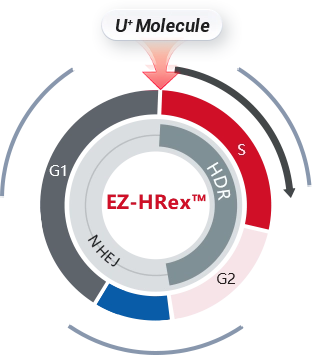
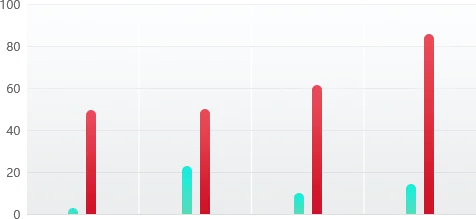
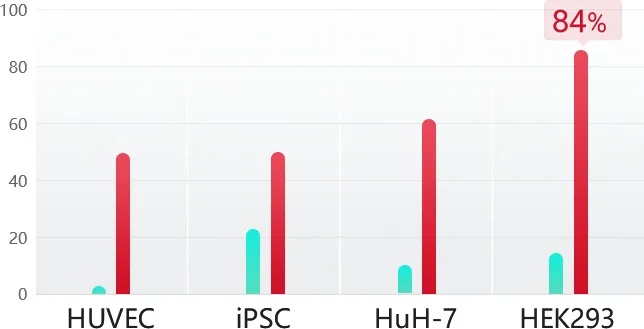
Case Study
HDR Efficiency (%)
48 %
HUVEC
50 %
iPSC
62 %
HuH-7
84 %
HEK293
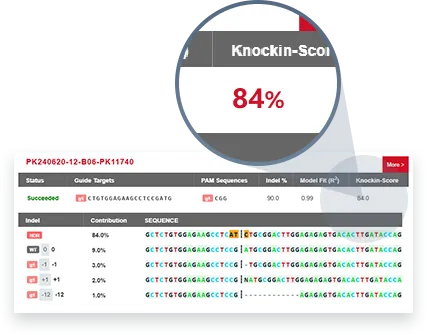
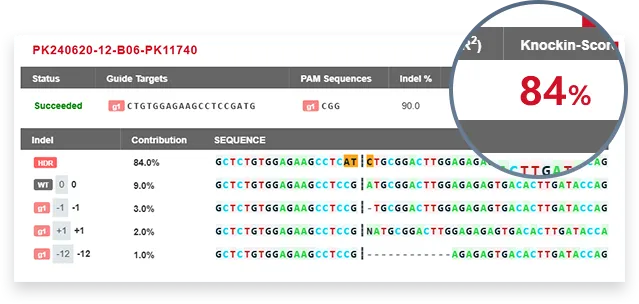
EZ-HRex™
Traditional Method

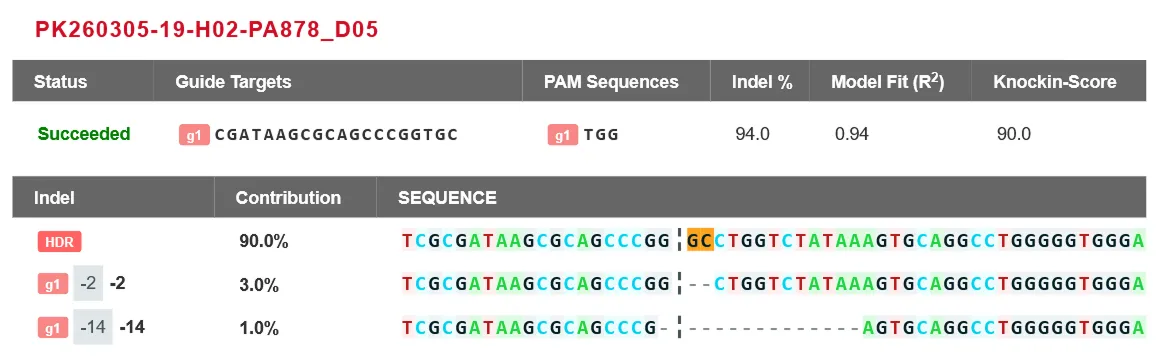
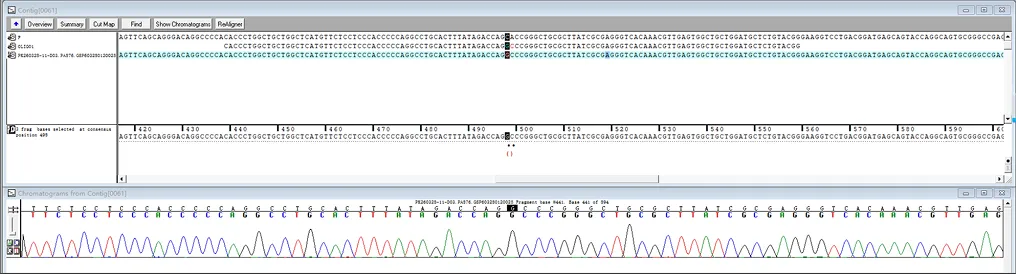
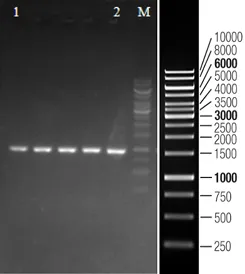
Featured Citations
A549 Cell MYC Gene - Lung Cancer
IF=20.8
Nature Metabolism
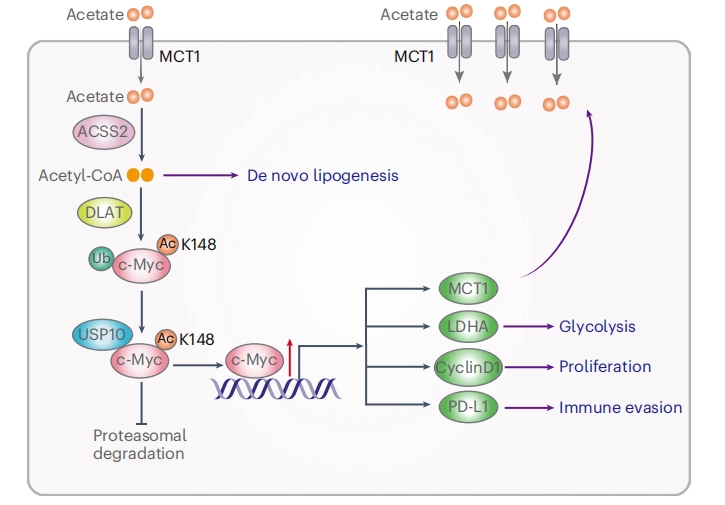
Acetate reprogrammes tumour metabolism and promotes PD-L1 expression and immune evasion by upregulating c-Myc
This study demonstrated that acetate is the most abundant short-chain fatty acid in human non-small cell lung cancer tissue, with increased acetate uptake as the tumors enriched. The study used Ubigene-constructed MYC (p.K148R) point mutant A549 cells and MYC (p.K148Q) point mutant A549 cells, simulating acetylated and non-acetylated c-Myc states, and investigated the impact of these changes on tumor cell behavior. View details
View Picture
FOXP1 gene in HEK293 cells - Tumor occurrence
IF=9.4
The EMBO Journal
FOXP1 phosphorylation antagonizes its O-GlcNAcylation in regulating ATR activation in response to replication stress
This study reveals a new mechanism by which the forkhead box (FOX) transcription factor FOXP1 promotes ATR-CHK1 activation under replication stress. This function does not rely on the transcriptional regulatory activity of FOXP1 but is instead co-regulated by its phosphorylation and glycosylation modifications. The study used the source cell lines from Genewell to construct the FOXP1 gene S396A and S396D point mutation HEK293 cell lines. View details
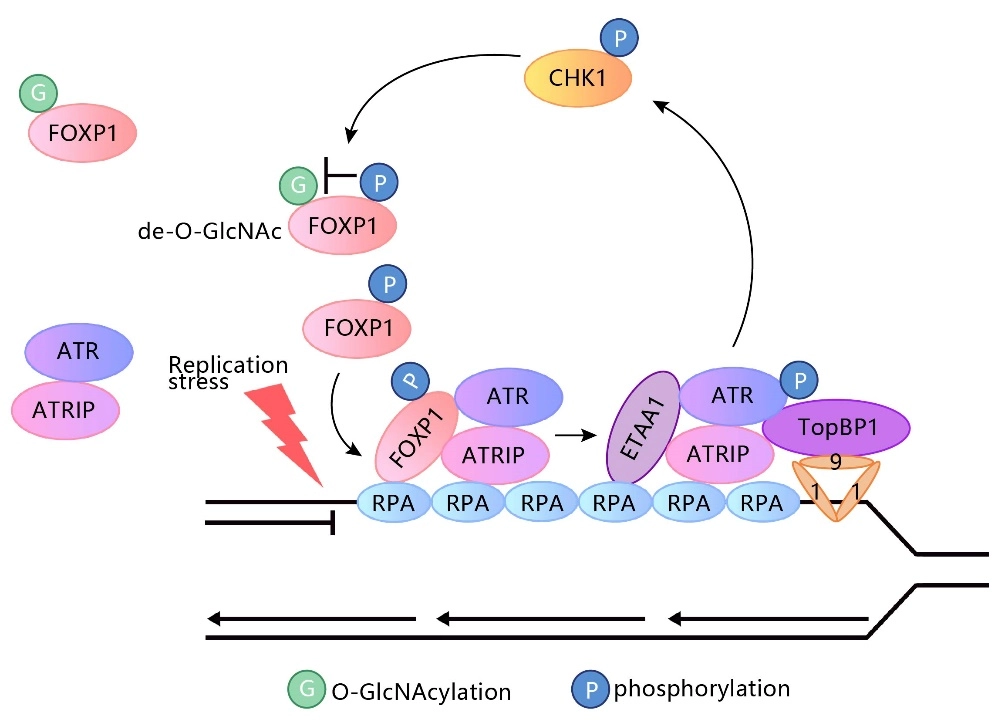
View Picture
FAQs
1. What is CRISPR point mutation?
CRISPR point mutation refers to using CRISPR/Cas9 technology to introduce a very specific, single-nucleotide change (A, T, C, or G) in the DNA sequence of a gene.
Instead of knocking out (deleting) a gene or inserting large DNA fragments (knock-in), point mutation precisely changes just one base pair. Scientists often use this to:
- Correct disease-causing mutations
- Create specific disease models
- Study the function of a particular amino acid in a protein
This is usually achieved by combining CRISPR/Cas9 with a carefully designed donor template (a short DNA sequence with the desired mutation) that the cell uses during repair.
2. What are the 4 types of point mutations?
- Substitution:
One base is replaced by another.
e.g. A changes to G. - Insertion:
One extra base is inserted into the DNA sequence.
e.g. An extra A is added between two bases. - Deletion:
One base is removed from the DNA sequence.
e.g. A single missing C. - Frameshift mutation (special case of insertion or deletion):
When an insertion or deletion changes the reading frame of the gene.
This can dramatically alter the resulting protein.
3. Are point mutations reversible?
Yes, especially with modern genetic technologies! Contact us now
There are 2 main lab strategies:
- CRISPR knock-in: Introduce a donor template with the correct base.
- Base editors: Use engineered CRISPR systems that can directly change one base to another without making a DNA break..
4. What are the applications of CRISPR point mutation?
- Disease modeling
- Gene function research
- Therapeutic correction
- Drug discovery and screening
- Agriculture and biotechnology