Frontier in Biotechnology | Up to 16.2-Fold Improvement in Editing Efficiency: AcrIIA27 Targets sgRNA to Overcome CRISPR Editing Bottlenecks

Introduction
The CRISPR–Cas system constitutes an adaptive immune mechanism in prokaryotes. In response, bacteriophages have evolved anti-CRISPR (Acr) proteins to counteract CRISPR-mediated defense. To date, multiple Acr proteins have been identified that inhibit CRISPR–Cas activity through diverse molecular mechanisms. However, inhibition mechanisms that directly target single-guide RNA (sgRNA) remain insufficiently characterized.A research team led by Prof. Zhiwei Huang at Harbin Institute of Technology has recently published a study in Nature Structural & Molecular Biology. Using cryo–electron microscopy (cryo-EM), the investigators resolved the structure of AcrIIA27 in complex with the Streptococcus pyogenes Cas9 (SpyCas9)–sgRNA ribonucleoprotein complex. Their findings elucidate the molecular mechanism by which AcrIIA27 targets sgRNA to suppress Cas9 activity.Building upon this mechanistic insight, the team developed a PTP RNA truncation strategy, which significantly enhances both the editing efficiency and specificity of CRISPR–Cas9 and TnpB-based genome editing systems, achieving improvements of up to 16.2-fold. This work provides a structural and mechanistic framework for rational engineering of next-generation genome editing platforms and offers new opportunities to overcome intrinsic limitations in CRISPR-mediated applications.

Research Background
The CRISPR–Cas system is widely distributed in bacteria and archaea, where it functions as an adaptive immune mechanism by recognizing and cleaving invading nucleic acids. To evade this defense, bacteriophages have evolved diverse anti-CRISPR (Acr) proteins. Reported mechanisms of Acr-mediated inhibition primarily include interference with CRISPR–Cas ribonucleoprotein (RNP) assembly, blockade of target nucleic acid binding, and suppression of nuclease activity. Broad-spectrum Acr proteins identified to date predominantly target conserved structural domains of Cas effector proteins. In contrast, no broadly acting Acr protein that exerts inhibition through direct targeting of single-guide RNA (sgRNA) has been reported. Meanwhile, CRISPR–Cas9 and emerging RNA-guided editors, such as TnpB, are constrained by suboptimal editing efficiency and off-target effects, limiting their translational and therapeutic potential. Systematic investigation of Acr inhibitory mechanisms may therefore provide fundamentally new strategies for the rational optimization of genome editing tools.
Research Objective
This study aimed to elucidate the molecular mechanism by which AcrIIA27 inhibits Cas9 activity through sgRNA targeting by resolving the cryo–electron microscopy (cryo-EM) structure of the AcrIIA27–SpyCas9–sgRNA complex. Furthermore, leveraging these mechanistic insights, the study sought to develop a generalizable strategy to enhance the editing efficiency of CRISPR–Cas and TnpB systems.
Research Strategy
- Structural Characterization: The SpyCas9–sgRNA–AcrIIA27 ternary complex was purified and subjected to cryo-EM analysis to determine its three-dimensional structure. The binding interface between AcrIIA27 and sgRNA was precisely mapped.
- Mechanistic Validation: Site-directed mutagenesis, biolayer interferometry (BLI), and in vitro DNA cleavage assays were performed to validate the roles of key AcrIIA27 residues in sgRNA binding and Cas9 inhibition. Structural comparisons further demonstrated that AcrIIA27 induces steric hindrance, thereby preventing substrate DNA binding.
- Broad-Spectrum Activity Assessment: Representative Cas9 homologs from type II-A, II-B, and II-C subtypes were evaluated to determine the inhibitory activity of AcrIIA27, establishing its broad-spectrum inhibitory profile.
- Genome Editing Optimization: Based on the sgRNA region bound by AcrIIA27 (designated PTP RNA), truncated sgRNA/reRNA variants were engineered. Their impact on editing efficiency and specificity of Cas9 and TnpB systems was evaluated in HEK293F cells.
- Mechanistic Elucidation of the Optimization Strategy: Integrating structural insights with cellular assays, the study proposes that PTP RNA truncation enhances genome editing efficiency by eliminating non-specific protein interactions and alleviating steric constraints, thereby facilitating more effective target DNA engagement.
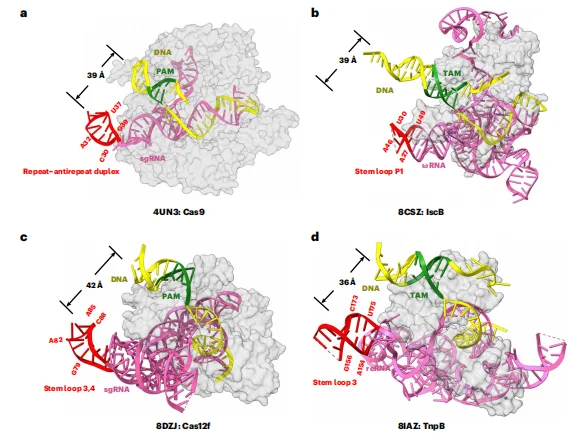
Figure 1 Application of the PTP RNA Truncation Strategy Across Multiple Genome Editing Platforms
Major Findings
1. Structural Features of the SpyCas9–sgRNA–AcrIIA27 Ternary Complex
Cryo–electron microscopy (cryo-EM) analysis revealed that Streptococcus pyogenes Cas9 (SpyCas9) adopts the canonical bilobed architecture consisting of a recognition (REC) lobe and a nuclease (NUC) lobe. AcrIIA27 binds to the SpyCas9–sgRNA ribonucleoprotein complex at a 1:1 stoichiometric ratio. Structurally, AcrIIA27 primarily engages the repeat–anti-repeat duplex region of the sgRNA (nucleotides G21–U45), while making limited contacts with the REC1 and WED domains of SpyCas9. This interaction mode highlights an RNA-centric inhibitory mechanism distinct from previously characterized Acr proteins that predominantly target conserved protein domains of Cas effectors. AcrIIA27 exhibits a unique structural fold composed of three β-strands, two α-helices, and three loop regions (loop1–loop3). A positively charged surface patch of AcrIIA27 inserts into the major groove of the sgRNA duplex, where electrostatic interactions and hydrogen bonding networks stabilize the protein–RNA interface.These structural features collectively define a previously uncharacterized mechanism of anti-CRISPR inhibition mediated through direct sgRNA engagement.
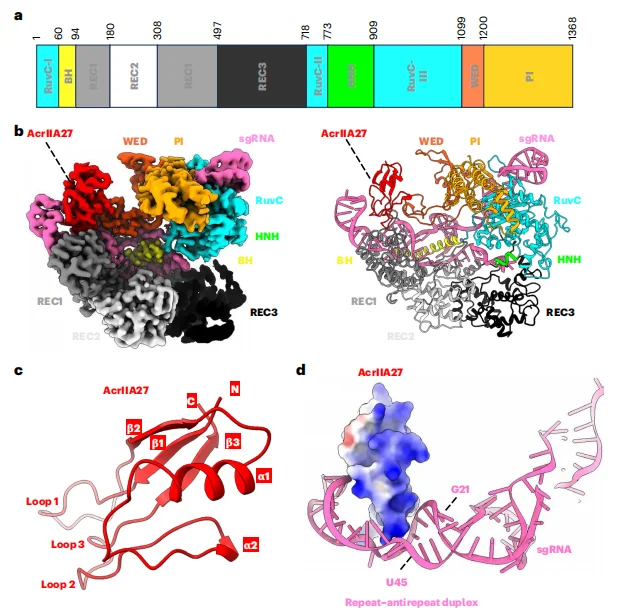
Figure 2. Cryo–EM Structure of the SpyCas9–sgRNA–AcrIIA27 Ternary Complex
2. AcrIIA27 Specifically Recognizes sgRNA Through Its Loop Regions
Structural analysis demonstrates that AcrIIA27 engages the sgRNA duplex primarily through its loop regions, which form a highly specific protein–RNA interaction interface.
(additional paragraph)Loop1-mediated interactions:
Positively charged residues within Loop1 (H17, R67, R71, and R72)
establish electrostatic interactions with the phosphate backbone of
sgRNA nucleotides such as U24 and A26. These charge–charge interactions
stabilize AcrIIA27 docking onto the repeat–anti-repeat duplex.
(additional paragraph) Loop2-mediated interactions:
Residues S48, K50, and W51 in Loop2 form hydrogen bonds with the
phosphate backbone of nucleotides U37 and A38, further reinforcing
binding specificity and affinity.
(additional paragraph) Biolayer interferometry (BLI) measurements revealed that wild-type AcrIIA27 binds the sgRNA-loaded Streptococcus pyogenes Cas9 complex with exceptionally high affinity (KD < 1 pM). In contrast, mutation of key Loop1 and Loop2 residues (e.g., R66A, R67A) resulted in more than a 10-fold reduction in binding affinity.Consistent with these findings, in vitro DNA cleavage assays showed that mutations at critical interface residues significantly impaired—or completely abolished—the inhibitory activity of AcrIIA27 against SpyCas9. Furthermore, bacteriophage plaque assays demonstrated that these loss-of-function mutants failed to restore the viability of T5 phage, confirming the functional importance of loop-mediated sgRNA recognition in AcrIIA27-dependent CRISPR inhibition.
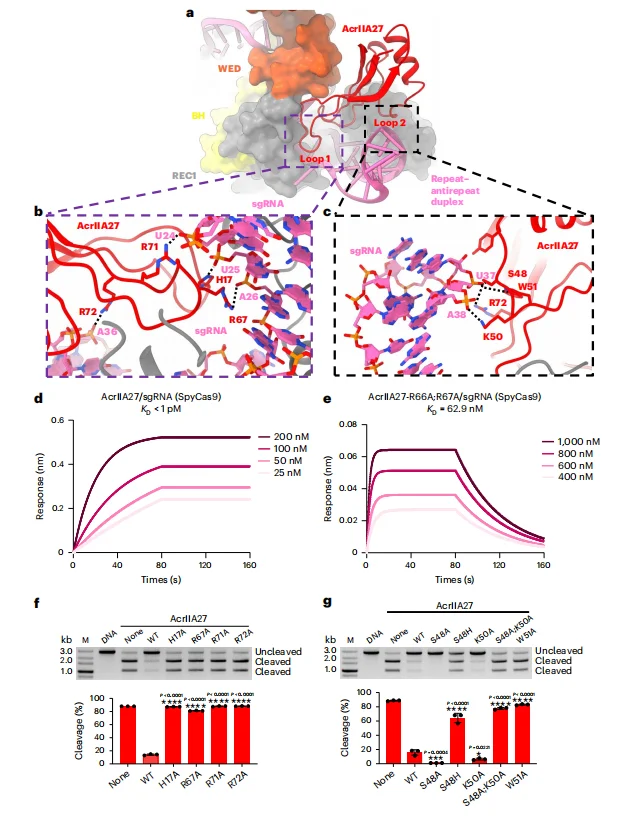
Figure 3. Structural Basis of sgRNA Recognition by AcrIIA27
3. Broad-Spectrum Inhibitory Activity of AcrIIA27 Across Cas9 Orthologs
Phylogenetic analysis demonstrated that the selected Cas9 homologs span multiple type II subfamilies, including:
- (additional paragraph) Type II-A: Streptococcus pyogenes Cas9, Staphylococcus aureus Cas9, Francisella novicida Cas9
- (additional paragraph) Type II-B: Francisella novicida Cas9
- (additional paragraph) Type II-C: Neisseria meningitidis Cas9, Neisseria meningitidis Cas9, Geobacillus stearothermophilus Cas9
(additional paragraph) In vitro DNA cleavage assays revealed that AcrIIA27 effectively inhibits the catalytic activity of type II-A and type II-C Cas9 orthologs. Notably, its inhibitory potency against SpyCas9 exceeded that of the subtype-specific inhibitor AcrIIA4. In contrast, AcrIIA27 exhibited no detectable inhibitory activity toward the type II-B effector FnCas9.Biolayer interferometry (BLI) further confirmed that AcrIIA27 binds with high affinity to sgRNA-loaded SauCas9 and Nme1Cas9 complexes. Mutation of critical interface residues significantly attenuated both binding affinity and inhibitory activity, indicating that the broad-spectrum suppression profile of AcrIIA27 is mechanistically dependent on its interaction with sgRNA rather than conserved protein domains.Collectively, these results establish AcrIIA27 as an RNA-targeting anti-CRISPR protein capable of broadly inhibiting type II-A and II-C Cas9 orthologs.
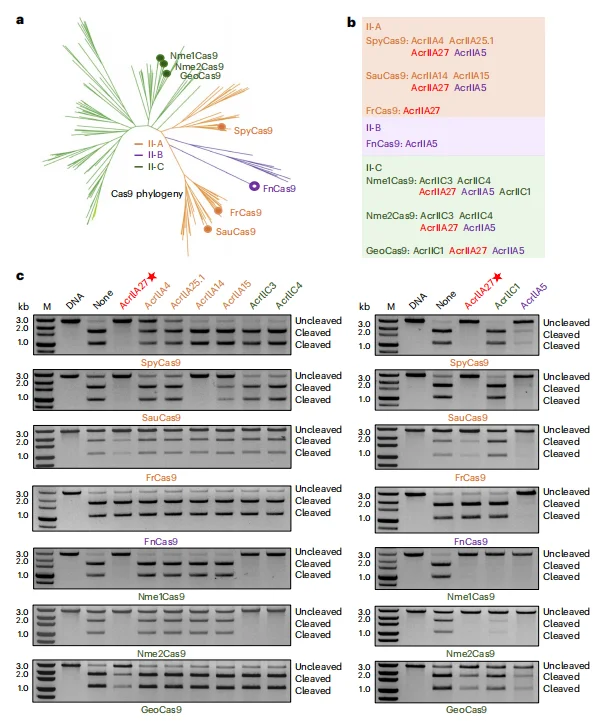
Figure 4. AcrIIA27 Inhibits Type II-A and Type II-C Cas9 Orthologs
4. AcrIIA27 Blocks Substrate DNA Binding to Cas9 via Steric Hindrance
Structural superposition of the AcrIIA27-bound complex with substrate DNA–loaded models revealed that the N54–T59 segment of AcrIIA27 spatially overlaps with the PAM-proximal region of the target DNA. This positional conflict indicates that AcrIIA27 sterically occludes the DNA binding channel required for productive target engagement.In vitro cleavage assays demonstrated that truncation of the N54–T59 segment completely abolished the inhibitory activity of AcrIIA27 against Streptococcus pyogenes Cas9. Importantly, this truncation did not impair its binding affinity to sgRNA, indicating that the loss of inhibition results specifically from disruption of steric interference rather than RNA binding deficiency.Conversely, truncation of the sgRNA stem–loop region (G29–C40) that serves as the AcrIIA27 interaction interface significantly attenuated AcrIIA27-mediated inhibition, while the intrinsic DNA cleavage activity of SpyCas9 remained unaffected. These results confirm that AcrIIA27 suppresses Cas9 function by occupying a spatial position that obstructs substrate DNA accommodation without compromising RNP assembly.Collectively, these findings establish steric hindrance of target DNA binding as the principal inhibitory mechanism of AcrIIA27.
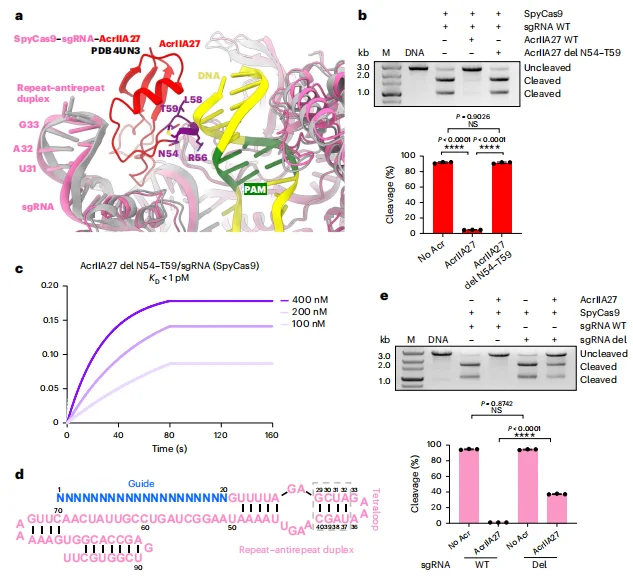
Figure 5. Mechanistic Basis of SpyCas9 Inhibition by AcrIIA27
5. PTP RNA Truncation Strategy Markedly Enhances Genome Editing Efficiency and Specificity
Application in SauCas9: For Staphylococcus aureus Cas9, truncation of the PAM-proximal PTP RNA region (U35–A60) within the sgRNA significantly improved genome editing performance. In HEK293F cells, editing efficiencies at multiple endogenous loci, including EMX1 and GRIN2B, increased by 1.2- to 2.1-fold compared with the full-length sgRNA configuration.
Application in IS607 TnpB Systems: In the IS607 TnpB system, truncation of the TAM-proximal PTP RNA region (corresponding to the MS1 site within stem-loop 3 of the reRNA) resulted in substantial efficiency gains. Editing efficiencies at loci such as EMX1 and DNMT1 improved by 1.5- to 6.1-fold.
Off-target analysis demonstrated that truncated PTP RNA designs not only enhanced on-target editing but also reduced off-target modification frequencies. Importantly, the increase in editing efficiency was independent of reRNA expression levels, indicating that the observed improvements arise from mechanistic optimization rather than elevated RNA abundance.This strategy was further validated in additional TnpB orthologs, including ISRin and ISAre variants, where editing efficiency improvements exceeded 16-fold at selected genomic loci.
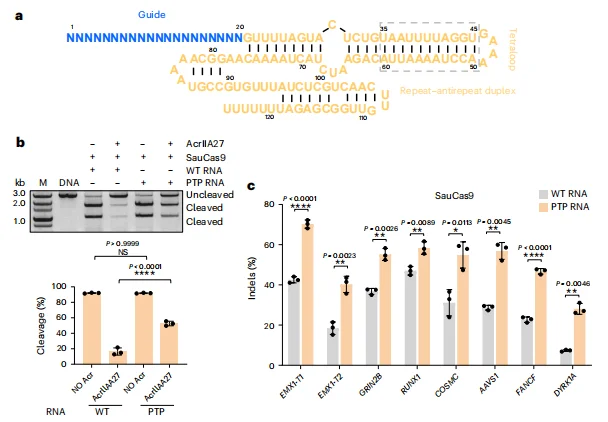
Figure 6. PAM-Proximal RNA Truncation Enhances Genome Editing Efficiency
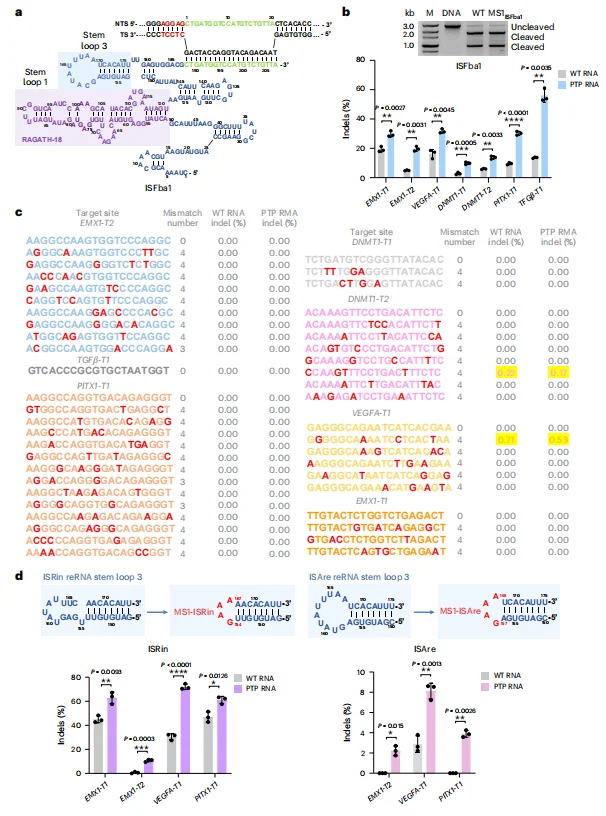
Figure 7. TAM-Proximal RNA Truncation Enhances Genome Editing Efficiency
Significance and Innovation
-
Theoretical Significance: This study represents the first structural and mechanistic characterization of an anti-CRISPR protein that inhibits Cas9 activity through direct targeting of the sgRNA phosphate backbone. By expanding the mechanistic classification of Acr proteins beyond protein-centric inhibition, it provides a conceptual framework for identifying novel Acr factors—particularly within RNA-binding protein families. Additionally, the study reveals that solvent-exposed regions of sgRNA (designated PTP RNA) are susceptible to non-specific interactions with positively charged intracellular proteins, which can impair editing efficiency. This finding offers new insight into intracellular regulatory influences on CRISPR activity and guide RNA performance.
-
Translational and Application Value: The proposed PTP RNA truncation strategy is broadly applicable and significantly enhances both efficiency and specificity of compact genome editing platforms, including CRISPR–Cas9 and TnpB systems—without requiring engineering of the nuclease protein itself. This simplifies optimization workflows and increases adaptability across editing modalities.Moreover, shortened sgRNA/reRNA architectures reduce overall payload size, improving compatibility with delivery vectors such as adeno-associated virus (AAV) and facilitating in vivo therapeutic genome editing applications.
-
Key Innovations: Challenges the traditional paradigm that anti-CRISPR proteins primarily target Cas effector proteins, by identifying the first broadly acting Acr that achieves inhibition through sgRNA engagement.Establishes sgRNA as a novel regulatory target for tunable genome editing control.Demonstrates a reverse-translation strategy: deriving an engineering solution (PTP RNA truncation) directly from structural insights into Acr-mediated inhibition, enabling rapid conversion of fundamental mechanistic discoveries into practical genome editing enhancements.
Summary
Through high-resolution cryo–electron microscopy and comprehensive biochemical validation, this study elucidates the molecular mechanism by which AcrIIA27 targets the PTP RNA region of sgRNA and inhibits Cas9 activity via steric blockade of substrate DNA binding.Mechanistically informed PTP RNA truncation effectively eliminates steric interference caused by non-specific protein interactions, thereby substantially enhancing genome editing efficiency and specificity of CRISPR–Cas9 and TnpB systems in human cells.Collectively, this work expands the functional repertoire of anti-CRISPR mechanisms and introduces a versatile, RNA-centric optimization strategy for developing next-generation high-efficiency and high-precision genome editing technologies.
Ubigene has remained committed to its core mission: “Make genome editing easier” Through continuous innovation and iterative optimization of both products and services, the company has successfully delivered more than 13,000 gene editing projects and has established a portfolio of over 11,000 engineered cell line products, including 8,000+ knockout (KO) cell lines. Leveraging proprietary technology platforms, Ubigene has achieved 10–20-fold higher genome editing efficiency compared with conventional methodologies. For precise point mutation engineering, the company has independently developed the EZ-HRex™ technology, which increases homology-directed repair (HDR) efficiency to as high as 84% post-transfection, substantially improving the success rate of precise genome modifications. (additional paragraph) Ubigene has provided high-quality genome editing services and products to more than 10,000 life science laboratories, pharmaceutical companies, and CROs worldwide, supporting research across basic science, drug discovery, and translational development.Ubigene will continue to advance its technology platforms, further enhance editing efficiency, and deliver high-performance, reliable genome engineering solutions to the global scientific community. (additional paragraph) For customized gene editing solutions, feel free to contact our team for consultation.
Contact us for more information >>>Reference
Yu L, Yin M, Zhu Y, Lu Z, Xiao B, Zhou F, Yu Y, Huang Z. An anti-CRISPR targets the sgRNA to block Cas9 and guides the design of enhanced genome editors. Nat Struct Mol Biol. 2026 Jan 19. doi: 10.1038/s41594-025-01741-z. Epub ahead of print. PMID: 41555077.



