Applications and Development Trends of CRISPR Library Screening in Drug Discovery

Challenges in Drug Discovery
Globally, the development of new drugs faces persistent challenges, including long development cycles, high R&D costs, and limited overall success rates. Statistics indicate that developing an innovative drug from early research to market approval typically requires 6–10 years or more, accompanied by substantial and sustained financial investment. However, the success rate of candidate drugs reaching the market remains low.
In the early stages of drug development, the discovery and functional validation of disease-related targets are particularly critical. These steps consume significant time and resources, as the biological relevance and druggability of a target directly affect the success of subsequent drug design and clinical development. Traditional target discovery strategies often rely on low-throughput or hypothesis-driven approaches, which are limited in systematically elucidating the causal relationships between gene function and disease phenotypes. Therefore, developing high-throughput and systematic technologies to enhance target identification and validation has become a key focus in modern drug discovery.
Value of CRISPR Library Screening
CRISPR library screening is a high-throughput functional genomics approach based on the CRISPR/Cas gene editing system. It allows systematic perturbation and functional evaluation of a large number of genes in a single experiment. Typically, this method involves constructing a pooled sgRNA library targeting either the whole genome or specific gene sets and introducing it into cell populations. Under uniform experimental conditions, the effects of perturbing different genes can be analyzed in parallel.
During screening, cells are exposed to specific selective pressures or experimental conditions, such as drug treatment, environmental stress, or phenotypic selection. Differences in proliferation, survival, or functional outcomes among cells carrying different sgRNAs are then quantified. High-throughput sequencing of sgRNA abundance before and after selection enables identification of genes that significantly impact cellular phenotypes, revealing their functional roles in biological processes or disease models.
Compared to phenotype- or molecular feature-based approaches, CRISPR library screening establishes a direct experimental causality between gene perturbation and phenotypic outcomes. This makes it a powerful tool for early-stage target discovery, functional gene identification, and mechanism-of-action studies, providing systematic, high-throughput support for drug development pipelines.
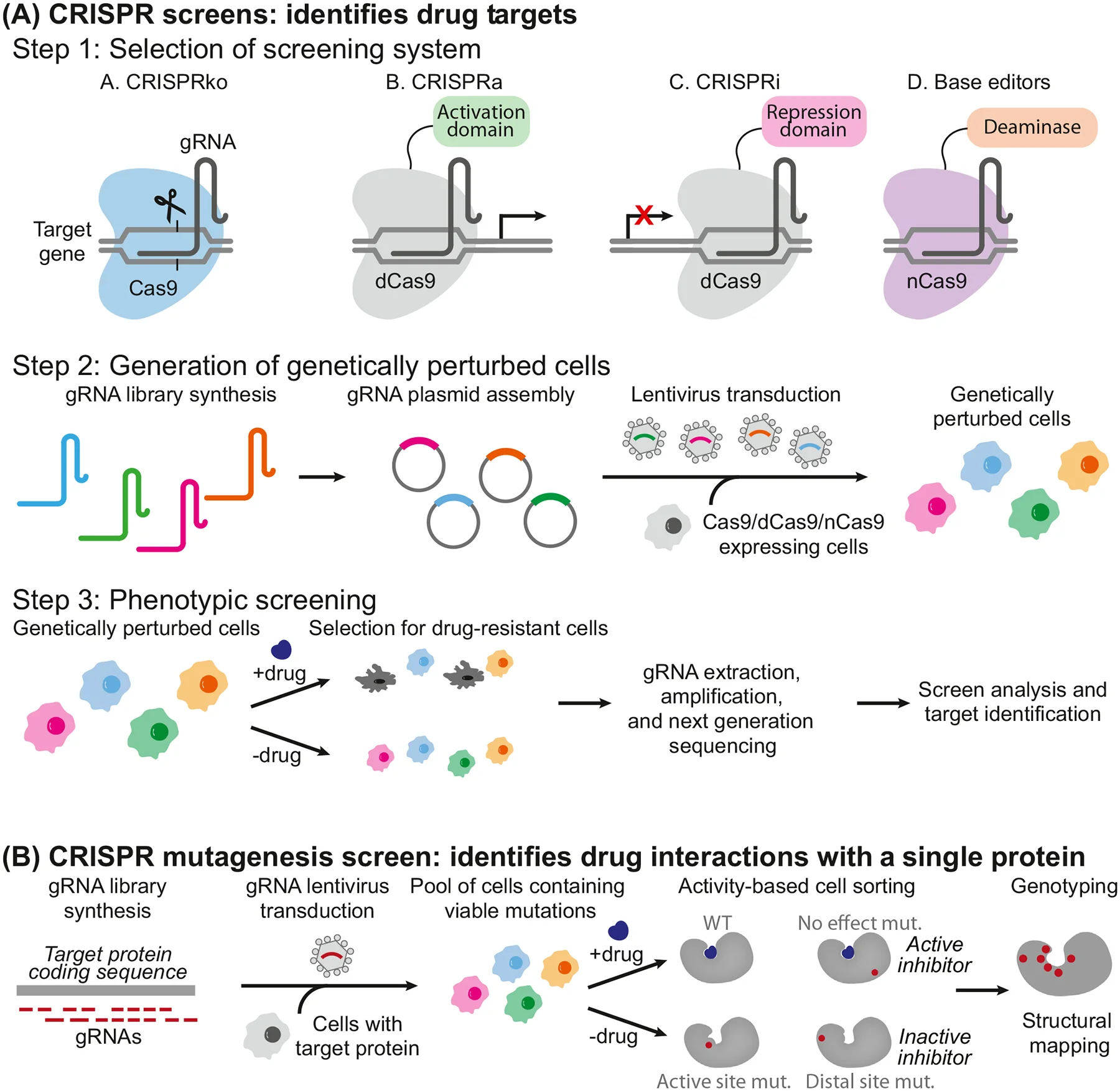
Figure 1. Schematic workflow of genome-wide CRISPR library screening. Genome-scale sgRNA libraries are constructed and introduced into cells via transfection or viral infection. Following application of drug treatment or other selective pressures, changes in sgRNA abundance before and after screening are analyzed by high-throughput sequencing to identify key genes.
Fundamental Principles of CRISPR Library Screening
CRISPR screening relies on the programmable gene-editing capability of the CRISPR-Cas system. By combining large-scale gene perturbation with phenotypic screening and sequencing analysis, it allows systematic functional annotation. The main CRISPR screening modes include:
- CRISPR-KO (Knockout): Uses nuclease-active Cas9 guided by sgRNA to introduce DNA double-strand breaks at target loci. Repair via Non-Homologous End Joining (NHEJ) often results in indels that disrupt reading frames, leading to gene loss-of-function. This mode is ideal for studying complete gene knockouts on cell phenotypes, biological processes, or disease models.
- CRISPRi (CRISPR interference): Employs catalytically inactive Cas9 (dCas9) fused to a transcriptional repressor domain (e.g., KRAB). Guided by sgRNA, it binds promoters or transcription start sites to block transcription without altering DNA sequences. CRISPRi allows stable and reversible gene expression knockdown, suitable for essential or dosage-sensitive genes.
- CRISPRa (CRISPR activation): Uses dCas9 fused with transcriptional activators (e.g., VP64, p65, Rta) to enhance endogenous gene expression at promoters or enhancers. CRISPRa enables gain-of-function studies, pathway activation analysis, and exploration of drug response mechanisms.
Together, CRISPR-KO, CRISPRi, and CRISPRa provide complementary ways to systematically perturb and evaluate gene function. Typical workflows include sgRNA library design and construction, viral packaging and cell infection, selective pressure application, high-throughput sequencing analysis, and identification of key gene targets.
Screening Strategies and Comparison
CRISPR library screening offers high flexibility in experimental design, allowing researchers to tailor strategies based on research goals, model systems, and desired phenotypes. Common screening dimensions include the following:
-
Screen Type: Positive vs. Negative Selection
- Positive selection enriches cells that gain a survival or proliferation advantage after gene perturbation under specific conditions, such as drug treatment. It is commonly used to identify genes conferring resistance or adaptability (e.g., drug-resistance genes).
- Negative selection (dropout screening) monitors the depletion of sgRNAs that impair cell growth, survival, or specific functions. It is ideal for discovering essential genes or targets that enhance drug sensitivity.
-
Screening System: In Vitro vs. In Vivo
- In vitro screening is performed in cultured cells, offering high throughput, controlled conditions, and suitability for large-scale target discovery and initial validation.
- In vivo screening uses animal models to evaluate gene function in a more physiologically relevant context, including tissue architecture, microenvironment, and cell-cell interactions. While more resource-intensive, it provides deeper insights for later-stage validation.
-
Editing Mode Selection: CRISPR-KO, CRISPRi, and CRISPRa
Researchers select the appropriate mode based on target gene characteristics:
- CRISPR-KO for complete loss-of-function studies (ideal for non-essential genes).
- CRISPRi for reversible knockdown (suitable for essential or dosage-sensitive genes).
- CRISPRa for gain-of-function studies (useful for pathway activation or drug response analysis).
Multi-mode or combinatorial screens are often employed for comprehensive results.
- Enrichment Methods and Phenotype Readout Strategies: Readouts can be flexibly designed, ranging from simple cell survival/proliferation-based population screening (via sgRNA abundance changes) to more advanced approaches. These include fluorescence reporters, surface markers with flow cytometry or magnetic sorting, and high-content imaging to capture complex phenotypes such as cell morphology, subcellular structures, or dynamic processes (e.g., stress granule formation).
Overall, the flexibility of CRISPR library screening enables systematic mapping of gene-phenotype relationships at genome scale. Careful consideration of research objectives, model systems, and technical feasibility during experimental design is essential to ensure biologically meaningful and reproducible outcomes.
Technical Features, Advantages, and Challenges
CRISPR library screening stands out as a powerful functional genomics tool for large-scale gene perturbation, offering significant value in target discovery and mechanism studies.
Key Advantages:
- High efficiency and strong signal-to-noise ratio: The CRISPR/Cas system generates stable and specific gene perturbations, making phenotypic changes easier to detect and quantify at the population level with reliable, reproducible results.
- Flexible experimental design: Multiple editing modes (CRISPR-KO for loss-of-function, CRISPRi for reversible knockdown, and CRISPRa for activation) can be combined with diverse selection pressures and models to address a wide range of research questions.
- Broad and systematic coverage: Libraries can target the whole genome, specific gene sets, or even regulatory regions (promoters and enhancers), enabling unbiased functional screening with minimal prior assumptions.
Challenges:
- Off-target effects and sgRNA quality: Even with high on-target precision, large-scale libraries require careful sgRNA design and multiple-guide validation to minimize false positives.
- Library construction and quality control: Genome-wide libraries demand high coverage (>99% recommended) and uniformity, involving complex synthesis, cloning, and validation steps that require robust technical platforms and expertise.
- Data analysis complexity: High-throughput sequencing data must account for sequencing depth, batch effects, and statistical rigor to identify meaningful hits.
- Limitations in complex phenotypes: Traditional readouts (e.g., survival or proliferation) may miss subtle or dynamic changes; high-content imaging or single-cell methods are increasingly needed.
Despite these challenges, successful implementation relies on thoughtful experimental design, high-quality platforms, and rigorous follow-up validation. When properly executed, CRISPR library screening delivers high-impact, scalable insights for drug discovery.
Applications in Drug Discovery
With the continuous advancement of technology and analytical methods, CRISPR library screening has been widely applied across various biomedical research fields, especially in gene function analysis and drug development. Key applications include:
- Cancer-Related Target Discovery: In oncology research, CRISPR library screening is commonly used to systematically identify genes critical for tumor cell growth, survival, and adaptability. For example, performing a genome-wide CRISPR-KO screen in melanoma cell lines carrying the BRAF V600E mutation using a GeCKO library under BRAF inhibitor Vemurafenib treatment can pinpoint candidate genes (e.g., NF1, MED12, CUL3) whose knockout confers drug resistance, providing experimental insight into resistance mechanisms.
- Drug Resistance and Mechanism Studies: CRISPR screens are widely used to study drug response mechanisms, identifying genes that influence sensitivity or resistance. By analyzing sgRNA enrichment or depletion under drug treatment, researchers can infer gene roles in drug response. CRISPRa-based screens can further explore the impact of gene upregulation on drug response, aiding in the dissection of drug action pathways and regulatory nodes.
- Synthetic Lethality Screening: Synthetic lethality screens are crucial for precision oncology. Dual-sgRNA libraries can simultaneously perturb two genes to identify gene pairs whose concurrent loss impairs cell viability. For instance, a library containing ~150,000 dual-sgRNA combinations can be used to screen multiple known cancer genes, revealing gene interactions with synthetic lethal relationships, providing new avenues for targeted therapy based on specific genetic backgrounds.
- Other Research Applications: Beyond oncology and drug development, CRISPR library screening is applied in infectious disease and immunity research, metabolic regulation studies, and signaling pathway analysis. For example, in viral infection models, CRISPR screens can identify host factors critical for pathogen entry or replication. In metabolism-related studies, large-scale gene perturbation can reveal key regulators of metabolic pathways. Across diverse biological processes, CRISPR screens serve as a versatile tool for functional genomics research.
Trends in Technology Integration
To overcome the limitations of traditional screens in phenotype capture and data depth, CRISPR library screening is increasingly integrated with advanced technologies to enable multi-dimensional functional analysis and target discovery. Current trends include:
- Integration with Single-Cell Omics: Combining CRISPR screens with single-cell RNA sequencing techniques (e.g., Perturb-seq, CROP-seq ) allows direct measurement of transcriptomic changes at the single-cell level. This enables simultaneous analysis of thousands to tens of thousands of gene perturbations in heterogeneous cell populations, revealing functional heterogeneity among cell types and states, and supporting detailed phenotypic resolution.
- Multi-Omics Integration: After CRISPR screens, integrating proteomics, metabolomics, and epigenomics can provide multi-layered validation and mechanistic insights. For instance, proteomic analysis of positive cells can construct gene function networks, while metabolomic profiling can elucidate gene impacts on metabolic regulation. This approach connects gene perturbations to molecular function, enhancing biological interpretation depth.
- High-Content Phenotypic Screening: High-content imaging technologies allow CRISPR screens to capture complex phenotypes not easily quantified by traditional readouts, such as changes in cell morphology, subcellular structures, or dynamic processes. CRaft-ID combines CRISPR-Cas9 screening with microraft arrays and high-resolution confocal imaging to monitor subcellular phenotypes, such as stress granule formation, at single-cell resolution, enabling high-throughput acquisition and quantification of phenotypic data.
Through these integrations, CRISPR library screening is evolving from single-phenotype observation to multi-dimensional, systematic, and quantitative functional analysis, providing richer data support and strategies for drug target discovery, mechanistic studies, and precision therapeutics.
Ubigene CRISPR Library Screening Platform
Ubigene leverages its proprietary CRISPR-iScreen™ technology platform to deliver end-to-end solutions for CRISPR library screening — from library design and construction to screening, sequencing, and data analysis. This integrated system supports gene function studies and drug target discovery with streamlined workflows.
Core Advantages of the CRISPR-iScreen™ Platform:
- High coverage and uniformity: Using high-efficiency competent cells and standardized cell pool preparation protocols, the platform achieves >99% library coverage with excellent uniformity (<10% deviation), providing a solid foundation for reliable screening results.
- Customizable multi-mode libraries: Supports CRISPR-KO, CRISPRi, and CRISPRa modes, allowing tailored libraries for knockout, transcriptional repression, or activation experiments to meet diverse research needs.
- Rich ready-to-use resources: Offers over 40 ready-made library plasmids and more than 400 standardized Cell Pool products, enabling quick experimental setup and accelerated research timelines.
- Versatile screening options: Compatible with both in vitro and in vivo models, and supports multiple selection pressures including drug treatment, passaging stress, viral infection, and FACS-based sorting.
- Advanced data analysis: The companion iScreenAnlys™, interactive platform provides user-friendly visualization, statistical analysis, and publication-ready outputs, enabling rapid interpretation from raw sequencing data to candidate targets.
By integrating library construction, cell handling, screening execution, high-throughput sequencing, and analysis into a unified system, the CRISPR-iScreen™ platform significantly reduces experimental complexity and shortens project timelines. It has been successfully applied in novel drug target discovery, functional gene studies, and mechanism validation, offering scalable technical support for academic and industry researchers.
Industry Development Trends
The application of CRISPR library screening in global drug discovery is growing rapidly. Numerous studies have demonstrated that this technology has been used by thousands of independent laboratories and R&D projects for gene function analysis and target identification. Industry reports indicate that as CRISPR screening becomes more widespread in academia and industry, the efficiency of novel drug target discovery continues to improve, gradually reaching scalable levels. Increasingly, pharmaceutical companies are integrating CRISPR library screening into early target validation workflows to support systematic candidate target selection and mechanistic studies. Key future development trends include:
- In vivo and patient-derived model applications: The technology is expanding from in vitro cell systems to in vivo animal models and patient-derived cells, allowing screening results to better reflect the effects of complex physiological environments, immune systems, and tissue microenvironments on gene function.
- AI- and data-driven library optimization: Combining artificial intelligence algorithms and big data analysis can optimize sgRNA design, library construction, and screening strategies, improving the accuracy and efficiency of target identification while reducing experimental costs and failure risks.
- Integration of CRISPR-derived technologies: Derived CRISPR techniques, such as base editing and prime editing, are gradually incorporated into screening systems, enabling non-knockout functional perturbations, precise gene expression regulation, and discovery of novel drug action sites.
- Deep integration with multi-omics and single-cell technologies: By combining CRISPR screening with single-cell omics, multi-omics, and high-content phenotypic analysis, researchers can achieve multi-layered, systematic functional insights, providing more comprehensive data support for complex disease models and precision drug development.
CRISPR library screening will continue to play a central role in drug target discovery, mechanistic research, and new drug development. The deep integration of CRISPR screening with single-cell, multi-omics, and intelligent data analysis represents a key direction for the future development of CRISPR technologies in the biopharmaceutical field.
Contact us to learn more>>>Reference
Modell AE, Lim D, Nguyen TM, Sreekanth V, Choudhary A. CRISPR-based therapeutics: current challenges and future applications. Trends Pharmacol Sci. 2022 Feb;43(2):151-161. doi: 10.1016/j.tips.2021.10.012. Epub 2021 Dec 21. PMID: 34952739; PMCID: PMC9726229.



