CRISPR Screening: KLHL6 as a Switch for T Cell Immunotherapy

Overview
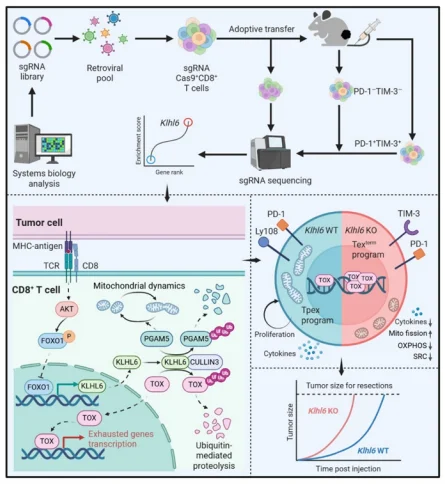
T cell exhaustion and mitochondrial dysfunction in tumor-infiltrating lymphocytes represent major barriers to effective cancer immunotherapy, while the role of proteostasis in regulating these processes remains poorly defined. Recently, a study by Li Guideng’s team published in Nature identified the E3 ubiquitin ligase KLHL6 as a dual negative regulator of both processes through integrative computational analysis and targeted in vivo CRISPR screening . Mechanistically, KLHL6 promotes ubiquitin-mediated degradation of TOX to suppress terminal T cell exhaustion, while maintaining mitochondrial homeostasis via regulation of the PGAM5–Drp1 axis. CRISPR screening enabled the discovery that T cell receptor (TCR) stimulation downregulates KLHL6 expression. Forced expression of KLHL6 significantly enhances the in vivo antitumor and antiviral efficacy and persistence of T cells, highlighting its potential as a clinically actionable target in cancer immunotherapy.

Research Background
In the tumor microenvironment or during chronic viral infections, CD8+ T cells undergo profound transcriptional and metabolic reprogramming due to persistent antigen exposure, ultimately leading to loss of effector function and sustained upregulation of inhibitory receptors. Previous studies focused on characterizing exhaustion states, but the molecular mechanisms—particularly the key regulatory nodes initiating exhaustion—remain insufficiently understood. Elucidating these mechanisms is critical for developing therapeutic strategies that sustain long-term T cell functionality.
Research Objectives
This study aims to identify and characterize the key molecular switch that governs the transition of CD8+ T cells from activation to exhaustion under persistent TCR stimulation. By integrating systems biology and functional genomics approaches, the authors sought to uncover core regulatory factors that simultaneously influence T cell exhaustion and mitochondrial metabolism, and to explore their potential applications in cancer immunotherapy.
Research Methods
Bioinformatics Analysis:
Transcriptomic datasets were leveraged to systematically characterize dynamic gene expression changes in T cells during acute versus chronic infection, enabling identification of candidate genes strongly associated with the exhaustion process.
In Vitro CRISPR Screening:
Lentiviral libraries targeting candidate genes were constructed and applied to primary CD8+ T cells for loss-of-function screening. Functional readouts, including cytokine secretion and inhibitory receptor expression, were used to identify key regulators of T cell exhaustion.
Molecular Biology and Biochemistry:
Co-immunoprecipitation (Co-IP) combined with mass spectrometry was used to identify substrate proteins of KLHL6; ubiquitination assays were performed to validate KLHL6-mediated degradation of its substrates.
Cellular Metabolism Analysis:
Mitochondrial respiratory function and dynamics (fission/fusion) were assessed using Seahorse extracellular flux analysis and confocal microscopy.
In Vivo Validation:
Adoptive T cell transfer models, including tumor models and chronic LCMV infection models, were used to evaluate the impact of KLHL6 modulation on T cell antitumor/antiviral function and exhaustion status.
Research Workflow
Differential gene screening:
Transcriptomic profiling of T cells was used to distinguish gene expression patterns between acute infection and chronic infection, identifying E3 ligase-associated proteostasis pathways as key regulators.
CRISPR library screening and validation:
In vivo CRISPR library screening combined with in vitro knockout and overexpression validation confirmed KLHL6 as a critical regulator of both T cell exhaustion and mitochondrial function.
Substrate identification and mechanistic analysis:
Direct substrates of KLHL6 were identified, revealing that KLHL6 targets TOX for degradation while also regulating the PGAM5–Drp1-mediated mitochondrial fission axis.
Metabolic–functional association analysis:
The study elucidated how KLHL6 deficiency-induced mitochondrial dysfunction further exacerbates T cell functional decline.
Translational relevance:
The therapeutic impact of enhancing KLHL6 expression was evaluated in tumor and chronic infection models, demonstrating its potential to improve immunotherapy outcomes.
Key Findings
1. KLHL6 is a dual negative regulator of T cell exhaustion and mitochondrial dysfunction
The authors refined transcriptomic variations from 136 CD8+ T cell RNA-seq samples across eight studies into two core modules to define the whole-transcriptome characteristics under chronic versus acute conditions. This confirmed the critical role of E3 ligase-mediated proteostasis in T cell exhaustion and mitochondrial maintenance. Parallel in vivo CRISPR screens targeting 78 exhaustion-associated and 133 mitochondria-associated E3 ligase genes consistently identified KLHL6 as a dual negative regulator.

Fig 1. KLHL6 is a dual negative regulator of T cell exhaustion and mitochondrial dysfunction.
2. KLHL6 deficiency drives terminal T cell exhaustion via TOX accumulation
Characterization of Klhl6−/− OT-I mice revealed that KLHL6 deficiency leads to reduced accumulation of CD8+ T cells in tumors, draining lymph nodes, and the spleen, alongside an increase in terminally exhausted T cells (PD-1+TIM-3+) and reduced secretion of TNF and IFNγ.
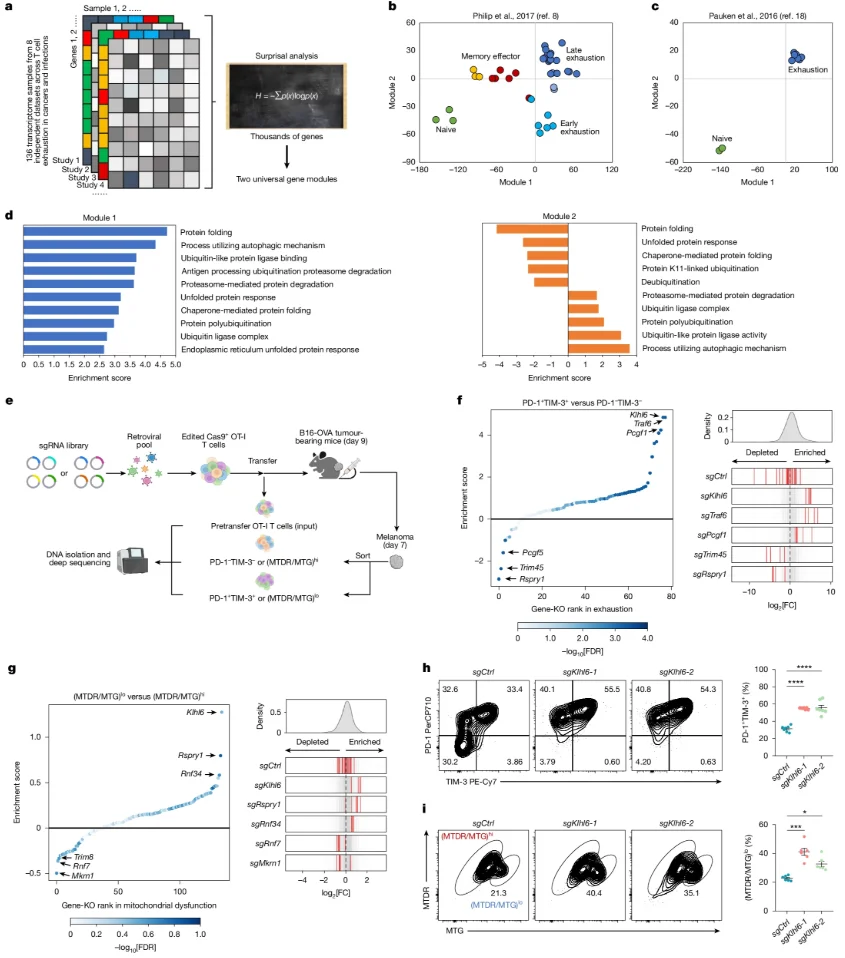
Fig 2. KLHL6 deficiency promotes T cell exhaustion and impairs function.
In contrast, B16-OVA mice treated with KLHL6-OE (overexpression) OT-I T cells showed increased T cell accumulation in the tumor and lymphoid organs, a lower proportion of terminally exhausted T cells, enhanced secretion of TNF and IFNγ, and prolonged survival. These data collectively demonstrate that the loss of KLHL6 drives CD8+ T cells toward exhaustion and dysfunction.
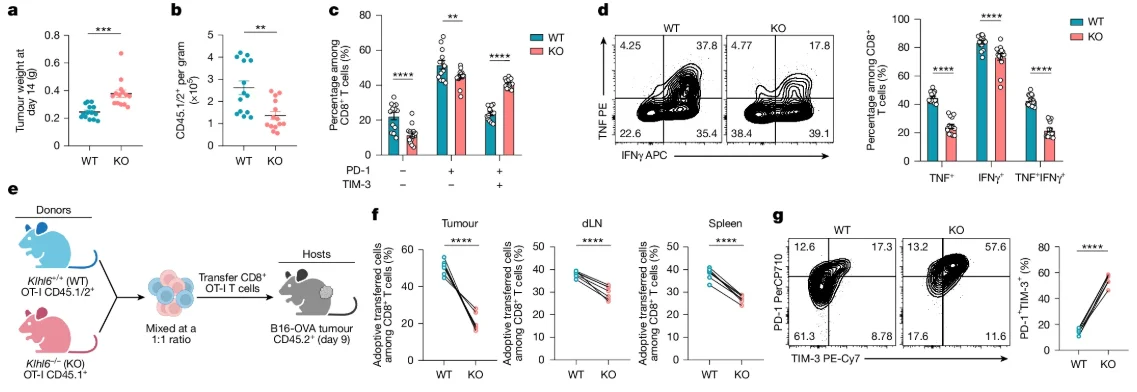
Fig 3. KLHL6 inhibits Tpex cell transition to terminal differentiation and enhances anti-tumor immunity.
3. TOX acts as a downstream target of KLHL6 to promote terminal differentiation
Further mechanistic study found that TOX is significantly enriched as a downstream substrate of KLHL6 during T cell exhaustion. KLHL6 modulation significantly regulates TOX protein levels through ubiquitination. While TOX is a known initiator of exhaustion, KLHL6-OE TILs showed reduced TOX and increased TCF-1 expression. Knocking down TOX in KLHL6-KO TILs reduced terminal exhaustion and increased precursor populations, partially inhibiting tumor progression. Pan-cancer scRNA-seq data from 316 patients further supported that KLHL6-mediated TOX regulation controls the transition from precursor to terminal exhaustion.
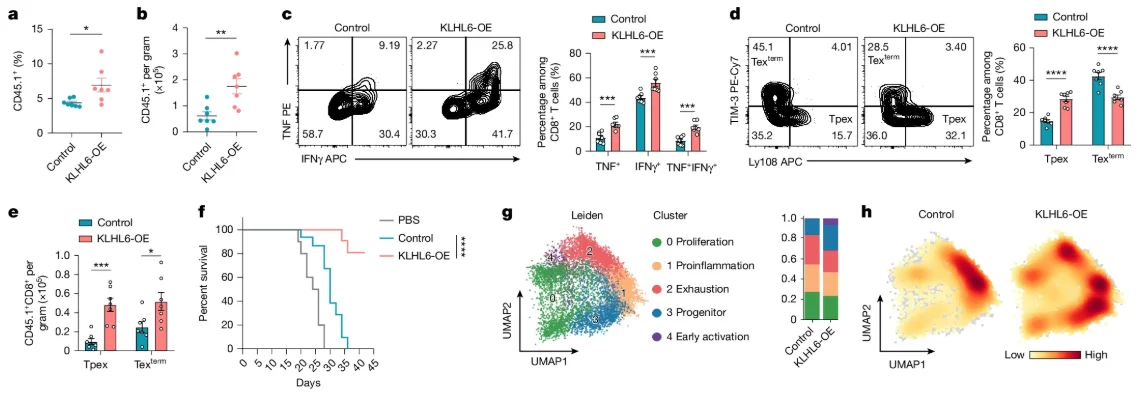
Fig 4. TOX as a downstream target of KLHL6 promotes terminal differentiation of Tex cells.
4. KLHL6 limits mitochondrial damage via the PGAM5-Drp1 axis
KLHL6-deficient T cells exhibit smaller mitochondria with shorter cristae, indicating a shift toward excessive fission. Knockdown of the downstream substrate PGAM5 restored fusion-related proteins (Mfn2, Opa1) and improved mitochondrial activity, oxygen consumption rate (OCR), and ATP production.
Overall, KLHL6 supports anti-tumor function by limiting TOX-driven exhaustion and PGAM5-mediated mitochondrial dysfunction. While single knockdown of TOX or PGAM5 only partially restored immunity, **dual knockdown resulted in almost complete rescue**, returning phenotypes and functions to wild-type levels. This underscores the coordinated regulation of KLHL6 across both transcriptional and metabolic pathways.
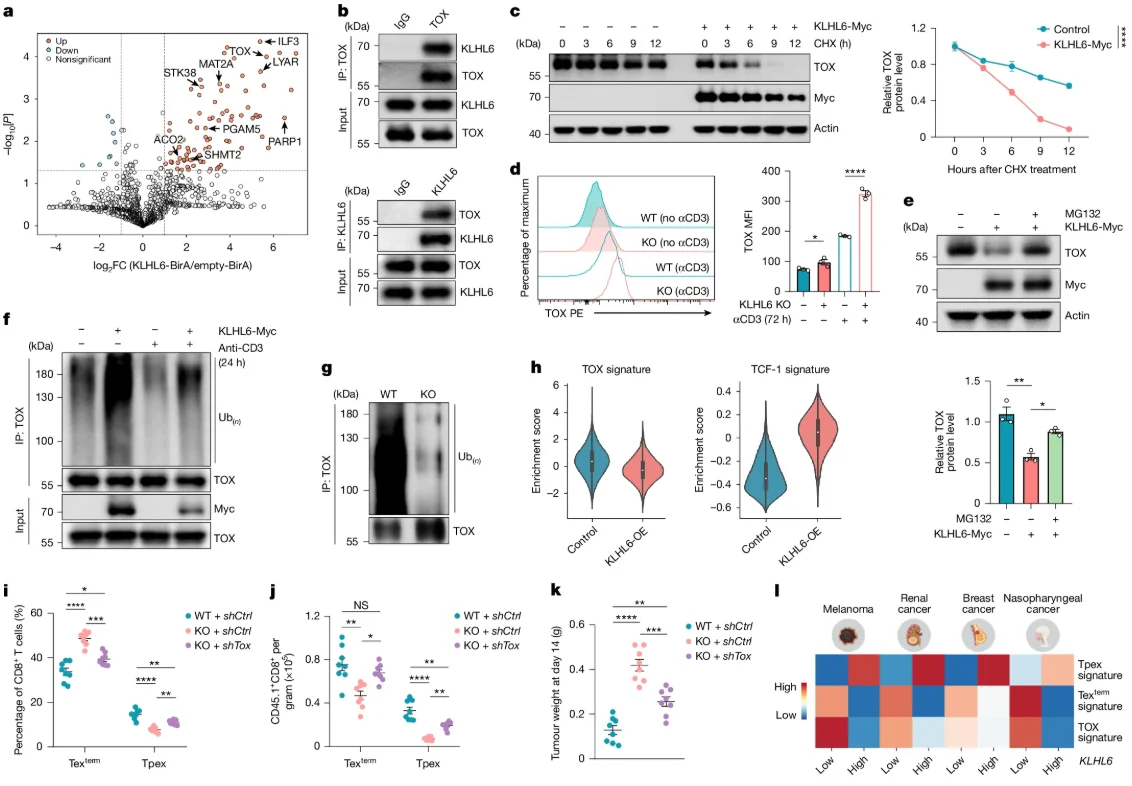
Fig 5. KLHL6 regulates T cell mitochondrial homeostasis via PGAM5 to modulate anti-tumor responses.
Summary
This study identifies the E3 ubiquitin ligase KLHL6 as a critical "gatekeeper" against T cell exhaustion. By targeting the exhaustion driver TOX for degradation and maintaining mitochondrial homeostasis, KLHL6 preserves T cell persistence. The discovery provides a highly promising therapeutic target for improving the efficacy of CAR-T and TCR-T therapies in solid tumors.
CRISPR-iScreen™ Libraries from Ubigene
CRISPR-iScreen™ is an innovative technology independently developed by Ubigene. Currently, Ubigene offers over 40 types of off-the-shelf CRISPR libraries and one-stop CRISPR functional screening services (in vivo/in vitro) to support your research.
Contact us for more details >>>


