CRISPR Screens Identify RARA/SPNS1 in Doxorubicin Cardiotoxicity

Title: CRISPR/Cas9 Screens Implicate RARA and SPNS1 in Doxorubicin Cardiotoxicity
Journal: JACC: CardioOncology IF=12.8
Research Institution: University of Pennsylvania.
Cell lines: Cas9-stable cardiomyocyte cell lines (HL-1cells, AC16 cells) and induced pluripotent stem cell-derived cardiomyocytes (iCMs)
Library Types: Genome-wide CRISPR/Cas9 knockout libraries (GeCKOv2, Brunello) and targeted CRISPR knockout libraries (targeting high-scoring hits, comprising 7,820 sgRNAs targeting 952 genes).
Screening Strategy: The study applied a dual-screening strategy (cell death screens + DOX accumulation screens) , using next-generation sequencing to analyze sgRNA abundance changes to identify genes regulating DOX-induced cell death, and flow cytometry to sort high-accumulation (top 15%) and low-accumulation (bottom 15%) populations to identify genes controlling DOX uptake, transport, or efflux.
Introduction
Anthracyclines such as doxorubicin (DOX) are widely used and vital anticancer drugs, particularly in childhood cancer and breast cancer, which are associated with 5-year survival rates of 85% or higher. However, it causes cardiotoxicity and heart failure, greatly limiting its clinical application, and the molecular etiology is poorly understood. Recently, a research team from the University of Pennsylvania published a landmark study in JACC: CardioOncology. They applied genome-wide and targeted CRISPR/Cas9 screening technologies with a functional genomics-based approach in an agnostic manner to systematically uncover genetic modifiers of DOX-induced cardiotoxicity, identifying for the first time retinoic acid receptor α (RARA) and sphingolipid transporter 1 (SPNS1) as key regulatory molecules. The study elucidated a dual mechanism: RARA activation reduces DOX toxicity by protecting mitochondrial function, whereas SPNS1 loss leads to abnormal DOX accumulation in lysosomes and exacerbates injury. This provides new targets and a technical framework for developing protective strategies against DOX cardiotoxicity.

Background
Clinical management of DOX cardiotoxicity has faced enormous challenges, and the understanding of its molecular etiology and genetic risk factors remains limited. Although some related genes and pathways have been identified, a large-scale, systematic mechanistic dissection is lacking. CRISPR screening, with its high specificity and genome-wide coverage, has become an ideal tool to uncover key regulatory genes in diseases. This study focused on DOX-induced cardiomyocyte injury and employed a dual-screening strategy (cell death regulators + DOX accumulation modifiers) to comprehensively identify genetic modifiers affecting DOX cardiotoxicity, elucidate their regulatory mechanisms, and provide a theoretical basis and potential targets for clinical protection.
Objectives
Adopting a functional genomics-based approach to systematically identify genetic modifiers of DOX-induced cardiotoxicity; Clarifying the molecular mechanisms by which RARA and SPNS1 regulate DOX cardiotoxicity; Validating the protective effects of RARA agonists and SPNS1-pathway targeting against DOX cardiotoxicity; and establishing a technical framework for the genetic regulation of DOX cardiotoxicity to support the discovery of therapeutic targets and genetic biomarkers.
Methods
Cell Models and Screening Systems
Constructing Cas9-stable cardiomyocyte cell lines (HL-1 cells, AC16 cells) and induced pluripotent stem cell-derived cardiomyocytes (iCMs) as the cell model for screening and validation; Genome-wide CRISPR/Cas9 knockout libraries (GeCKOv2, Brunello) and targeted CRISPR knockout libraries (targeting high-scoring hits, comprising 7,820 sgRNAs targeting 952 genes) were performed in screening.
Dual Screening Strategy
- Cell death screens: Library-transduced cardiomyocytes were divided into DOX-treated (1 μmol/L, 72 h, LD90 dose) and DMSO control groups; sgRNA abundance changes were analyzed by next-generation sequencing to identify genes regulating DOX-induced cell death;
- DOX accumulation screens: AC16 cells transduced with the library were treated with DOX (1 μmol/L, 24 h) and then sorted by flow cytometry into high-accumulation (top 15%) and low-accumulation (bottom 15%) populations; sequencing identified genes controlling DOX uptake, transport, or efflux.
Functional Validation and Mechanistic Illumination
RARA and SPNS1 knockout cell lines (HL-1, AC16, iPSCs) were generated via CRISPR-Cas9. Cell viability was measured by alamarBlue assay; DOX accumulation was quantified by flow cytometry and fluorescence microscopy. RNA-Seq was used to analyze gene expression changes, focusing on mitochondria-related pathways. Lysosomal function, autophagic flux, and DNA damage markers (γ-H2AX) were assessed using LysoTracker staining, immunofluorescence, and Western blot. Spatiotemporal dynamics of DOX accumulation were observed by time-resolved confocal microscopy.
Pharmacological Validation
The protective effect of the RARA agonist tamibarotene (TBT) against DOX toxicity was evaluated in iCMs. Autophagy was activated by rapamycin or Torin1 to test its impact on DOX accumulation in SPNS1-deficient cells. DOX accumulation and toxicity were also examined under nutrient-deprivation conditions (HBSS medium).
Workflow
- Genome-wide screens: Genome-wide CRISPR screens for DOX-induced cell death and DOX accumulation were performed in HL-1 and AC16 cells to initially identify candidate regulatory genes;
- Targeted validation screen: A targeted CRISPR screen was performed in iCMs focusing on high-confidence targets, confirming the core regulatory roles of RARA and SPNS1;
- Mechanism illumination: Elucidating the molecular mechanisms of RARA in regulating mitochondrial function and SPNS1 in regulating lysosomal homeostasis and DOX handling;
- Pharmacological validation: Protective strategies such as RARA agonism and nutrient deprivation were tested against DOX cardiotoxicity.
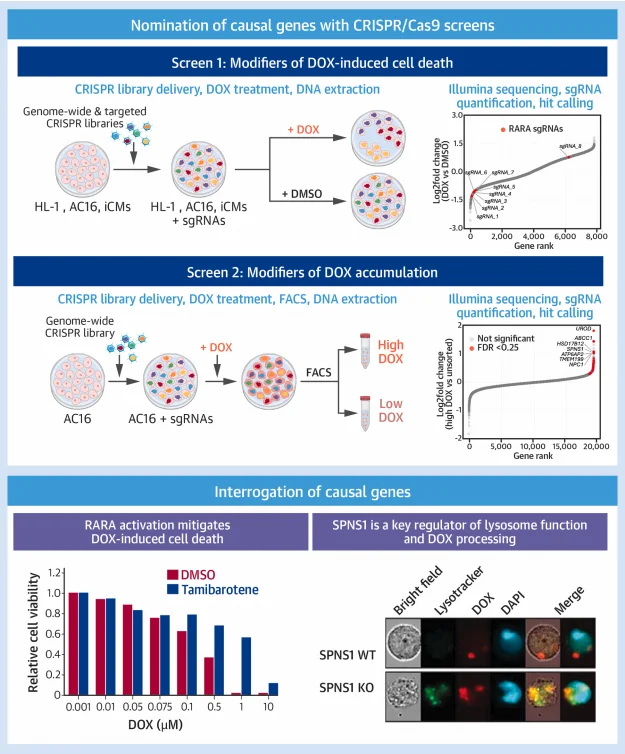
Figure 1. Screening strategy of the study
Results
1. Genome-wide CRISPR screens identify Modifiers of DOX-induced cell death
In HL-1 cells, 36 desensitizing genes (sgRNA enrichment) and 87 sensitizing genes (sgRNA depletion) were identified. Loss of Top2A, the primary DOX target, was the strongest desensitizing factor (log2FC=2.1, FDR<0.001), while loss of the DOX efflux transporter Abcb1b was the strongest sensitizing factor (log2FC=-1.9, FDR<0.001). In AC16 cells, 69 desensitizing and 25 sensitizing genes were identified. GO analysis showed that these genes were significantly enriched in pathways relevant to DOX cardiotoxicity, such as DNA damage response and apoptosis. Parallel screening in MCF7 cells ruled out some cancer-cell-specific modifiers, ensuring the relevance of candidate genes to DOX cardiotoxicity in cardiomyocytes.
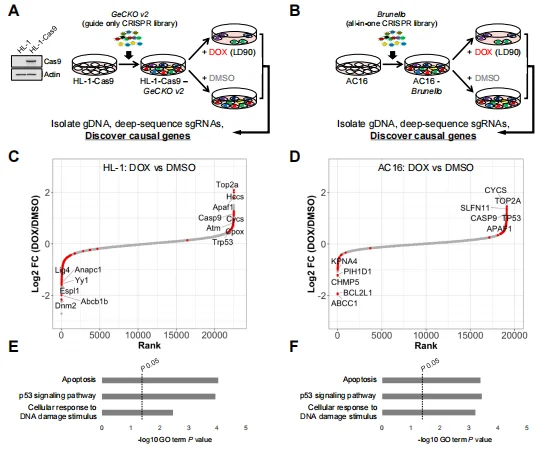
Figure 2. Genome-wide CRISPR/Cas9 knockout screens identify modifiers of DOX cytotoxicity
2. Targeted CRISPR screen validates RARA as a key regulator of DOX toxicity
In iCMs, RARA was the most significant sensitizing gene, with its targeting sgRNAs significantly depleted in the DOX-treated group. CRISPR-mediated RARA knockout (RARA-KO) iCMs showed markedly increased sensitivity to DOX, with significantly lower cell viability than wild-type (WT) iCMs at equivalent doses. Pretreatment with the RARA agonist tamibarotene (TBT) significantly reduced DOX-induced iCM death without affecting DOX cytotoxicity in MCF7 cancer cells; TBT also alleviated DOX-induced myofibrillar disarray in iCMs, confirming a cardiomyocyte-specific protective effect of RARA activation.
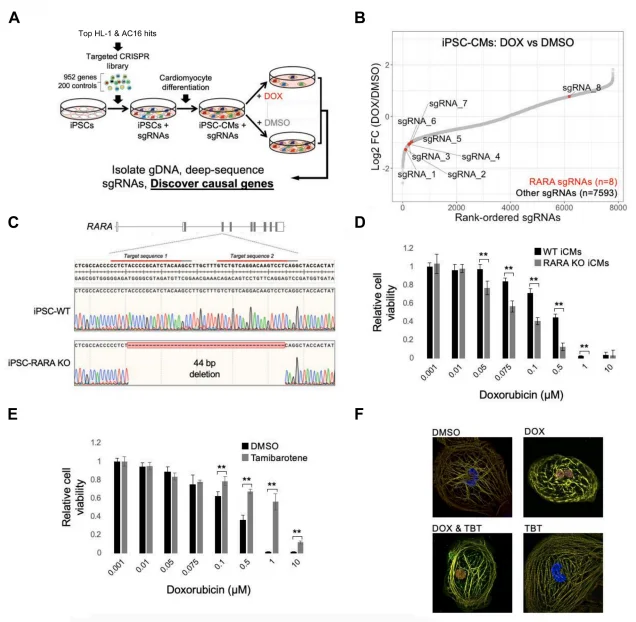
Figure 3. Validation of RARA status as a determinant of DOX cardiotoxicity
3. RARA activation reduces DOX toxicity by protecting mitochondrial gene expression
RNA-Seq analysis showed that DOX treatment significantly downregulated the expression of genes encoding mitochondrially localized proteins in iCMs, whereas combined TBT and DOX treatment reversed this inhibitory effect. Principal component analysis and volcano plots indicated that TBT treatment specifically modulated DOX-induced transcriptomic changes, especially in mitochondria-related pathways, suggesting that RARA activation alleviates toxicity by buffering DOX-mediated suppression of mitochondrial gene expression and improving cardiomyocyte metabolic function.
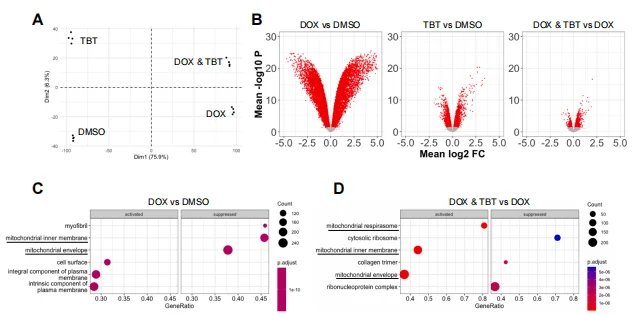
Figure 4. TBT Mitigates DOX-Mediated Suppression of Mitochondrial Gene Expression
4. Genome-wide screen identifies modifiers of DOX accumulation and highlights lysosomal function
The DOX accumulation screen showed that DOX fluorescence intensity in AC16 cells increased dose-dependently, and ABCC1 (a known DOX efflux pump) knockout cells accumulated significantly more DOX than WT cells, validating the screening system. The screen identified 32 genes associated with low DOX accumulation and 23 genes associated with high DOX accumulation. GO analysis revealed that high-accumulation-associated genes were primarily localized to or regulated endosome/lysosome function. SPNS1 was a core hit in the high-accumulation group (log2FC=1.1, FDR<0.001), and its function had not previously been linked to DOX handling.
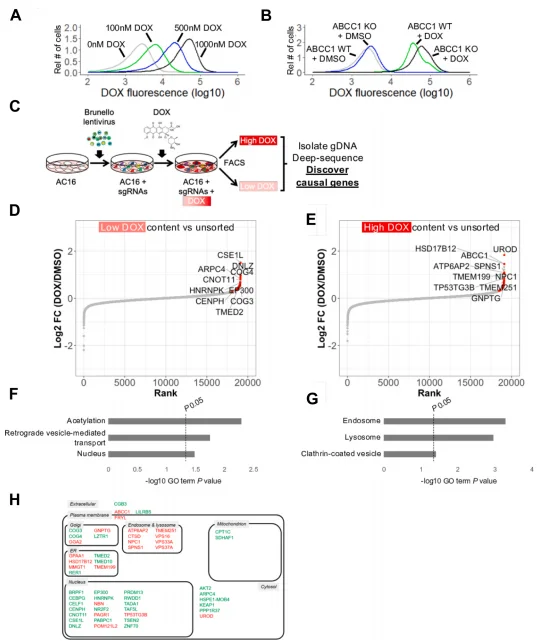
Figure 5 Discovery of Factors Regulating DOX Accumulation via Genome-Wide CRISPR/Cas9 KO Screens in AC16 Human Cardiomyocytes
5. SPNS1 loss leads to abnormal DOX accumulation and exacerbates cardiomyocyte injury
In SPNS1-KO AC16 cells and iCMs, DOX accumulation was significantly higher than in WT cells, with faster accumulation kinetics, and a large amount of DOX localized to perinuclear bodies (lysosomes). LysoTracker staining confirmed increased lysosome number in SPNS1-KO cells and significant co-localization with DOX. SPNS1 deficiency also led to abnormal autophagic flux (increased LC3 puncta, accumulation of LC3-II) and exacerbated DNA damage (elevated γ-H2AX expression). In SPNS1-KO cells, activating autophagy with rapamycin or Torin1 further increased DOX accumulation, and SPNS1-KO iCMs were significantly more sensitive to DOX toxicity than WT cells, confirming that SPNS1-mediated coupling of lysosomal homeostasis and autophagy a key modifier of DOX uptake, accumulation, and toxicity.
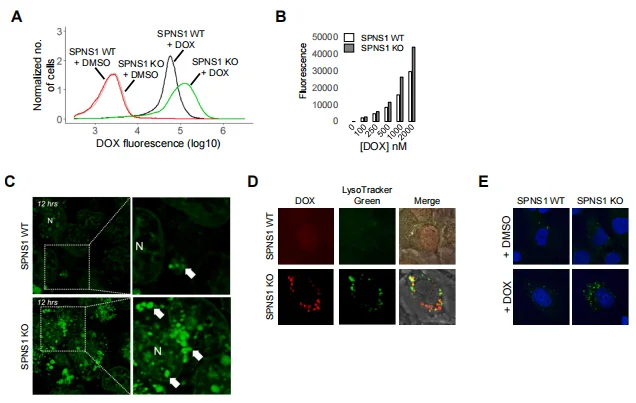
Figure 6. Validation of SPNS1 as a Modifier of DOX Accumulation
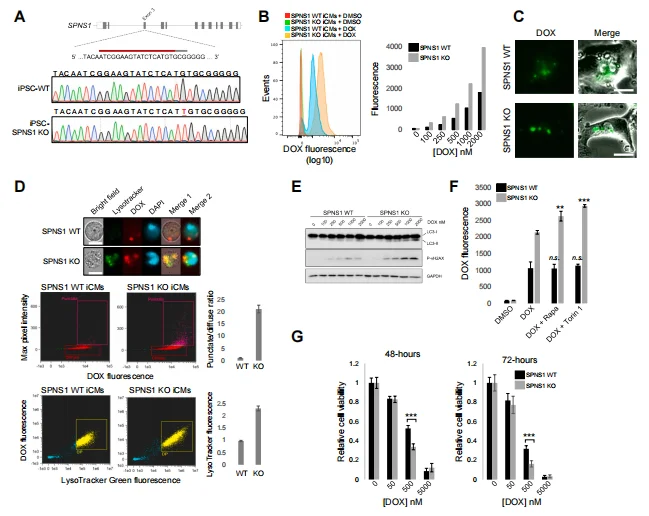
Figure 7. SPNS1 Deficiency Exacerbates DOX Accumulation and Toxicity in iPSC-Derived Cardiomyocytes
6. Ribosome loss-of-function and nutrient depletion reduce DOX accumulation and toxicity
Secondary analysis of screen data revealed that loss of genes involved in ribosome biogenesis and function was strongly associated with reduced DOX accumulation and increased survival. Under nutrient-deprivation conditions (HBSS medium), iCMs showed significantly reduced DOX accumulation and decreased DOX-induced cell death; this effect was reversed by nutrient replenishment, suggesting that nutrient availability regulates DOX accumulation and toxicity through a non-mTORC pathway.
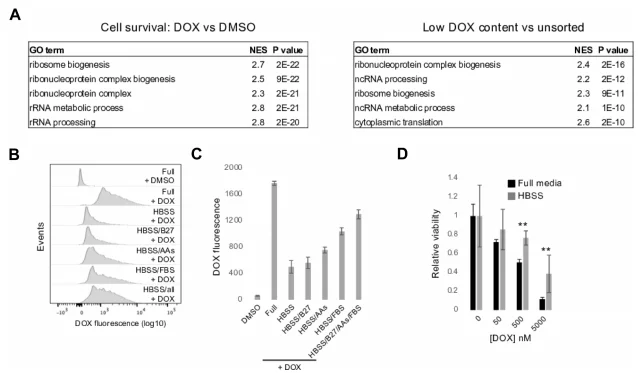
Figure 8. Loss of Ribosome Genes and Nutrient Deprivation Reduce DOX Accumulation and Toxicity
Significance and Innovations
Mechanistic Innovation
For the first time, a dual CRISPR screening system systematically identified two classes of modifiers of DOX cardiotoxicity, elucidating two independent and critical pathways—"RARA–mitochondrial function" and "SPNS1–lysosomal homeostasis–autophagy"—breaking the perception of a single mechanism and revealing the important influence of genetic background on DOX response.
Target and Strategy Innovation
Identified RARA as a protective target against DOX cardiotoxicity, with its agonist tamibarotene specifically protecting cardiomyocytes. Defined SPNS1 as a modifier of lysosomal function, providing a basis for developing protective strategies targeting lysosomal pathways. Discovered that nutrient deprivation reduces DOX accumulation, offering a new approach for optimizing clinical dosing regimens.
Technical Framework Innovation
Established a complete functional genomics workflow of "genome-wide screen → targeted validation → mechanistic illumination → pharmacological validation," providing a reproducible technical paradigm for studying the genetic regulation of other chemotherapy-induced toxicities.
Conclusions
This study systematically identified RARA and SPNS1 as key modifiers of DOX cardiotoxicity through genome-wide and targeted CRISPR screens, elucidating a dual regulatory mechanism: RARA loss sensitizes cardiomyocytes to DOX, while RARA activation reduces toxicity by protecting mitochondrial gene expression; SPNS1 loss leads to lysosomal dyshomeostasis, abnormal DOX accumulation, suppressed autophagy, and exacerbated DNA damage and cell death. Additionally, ribosome loss-of-function and nutrient depletion reduce DOX accumulation, providing complementary avenues for protective strategies. This work not only deepens our understanding of the molecular mechanisms of DOX cardiotoxicity but also identifies potential therapeutic targets (RARA, SPNS1) and feasible protective strategies (RARA agonists, nutrient modulation), providing important theoretical and experimental bases for improving the safety of DOX therapy in clinical practice.
Ubigene CRISPR-iScreen™ Library
CRISPR-iScreen™ is an innovative technology independently developed by Ubigene, designed to achieve efficient CRISPR screening. Currently, Ubigene offers over 40 CRISPR libraries in stock, covering genome-wide knockout/ inhibition/activation libraries for humans, mice, green monkeys, pigs, and other species. Additionally, there are CRISPR knockout/ inhibition/activation sub-libraries targeting genes related to kinases, the cell cycle, membrane proteins, and metabolism, helping you easily identify targets of interest! Ubigene currently has over 150 library virus stocks and over 600 library Cell Pools, and also provides one-stop CRISPR library (in vitro/in vivo) functional screening services to readily support your research.
Ubigene New Year Special Offer: CRISPR library NGS sequencing (including analysis) as low as $1600 , library data analysis as low as $500 , free use of iScreenAnlys™ library analysis platform, free choice of Drug-Z/MAGeCK-MLE/MAGeCK-RRA algorithms, one-click analysis of screening data. Contact us to learn more>>>
Reference
McDermott-Roe C, Lv W, Shao Y, Hoshino A, Arany Z, Musunuru K. CRISPR/Cas9 Screens Implicate RARA and SPNS1 in Doxorubicin Cardiotoxicity. JACC CardioOncol. 2026 Feb;8(1):48-61. doi: 10.1016/j.jaccao.2026.01.003. PMID: 41705753; PMCID: PMC12922622.



