[Research Highlight] CRISPR-B™ technology helps reveal the relation between Salmonella and Fe | Ubigene
Salmonellosis (Salmonella infection) is a common bacterial disease caused by various types of Salmonella in humans, poultry as well as wild birds and small mammals. Food can be contaminated by Salmonella in the feces of infected humans or other animals, and food poisoning can occur in humans. According to statistics in bacterial food poisoning in countries around the world, food poisoning caused by Salmonella is often ranked at the top. Therefore, investigating the mechanism of survival adaptation of Salmonella within the host is important for treatment after Salmonella infection.
Like other Enterobacteriaceae, Salmonella also requires the cofactor Fe to participate in multiple vital life processes such as nitrogen fixation, DNA synthesis, and damage repair. Fe in nature, however, is difficult to be directly utilized by Salmonella, so obtaining sufficient Fe from host cells by Salmonella is essential for their colonization and survival. But the host can limit its uptake of Fe by nutritional immunity, thus Salmonella evolved a catechol siderophore with high iron affinity, Enterobactin (Ent), to combat the host's nutritional immunity. Paradoxically, high concentrations of Fe produce a Fenton reaction that produces large amounts of hydroxyl radicals, leading to the death of Salmonella. Eventually, during the course of the relationship between Salmonella and its host, Salmonella have evolved iron homeostatic mechanism that allow it to acquire the required iron while avoiding toxic responses caused by excess iron.
Focusing on this iron homeostatic mechanism, Bingqing Li's team from Institute of Basic Medicine, Shandong First Medical University & Shandong Academy of Medical Sciences, studied Salmonella in the aspect of the post-translational modification of proteins and found that Salmonella, through Fe metabolism relating gene YdiU, could directly modify the highly conserved critical iron homeostasis regulator, Fur, by UMPylation, which abolishes the Fe uptake inhibition ability of fur, ultimately facilitating the uptake of Fe in Salmonella when lack of Fe. And the H118 site was also found to be the major UMPylation site of Fur, which has great significance for the effective regulation of iron homeostasis mechanism in Salmonella.
As a first step, researchers mimicked the Fe deficient environment, and used qPCR and Western blot to test the expression amount of YdiU at mRNA and protein levels under different dipyridyl concentrations as well as under 600μM dipyridyl condition for different times (Figure 1A-D). The results showed that the expression of YdiU could be effectively induced by dipyridyl, indicating that YdiU can respond to metal starvation signals. And by observing the dipyridyl or normal LB medium treated the YdiU knockout strain (ΔYdiU) and the wild-type strain (WT), it was further indicated through bacterial survival that YdiU can be effectively induced by metal deficiency signals, while the iron-add-back experiment (Figure 1H) can illustrate the involvement of YdiU in the metabolic pathway of Fe.
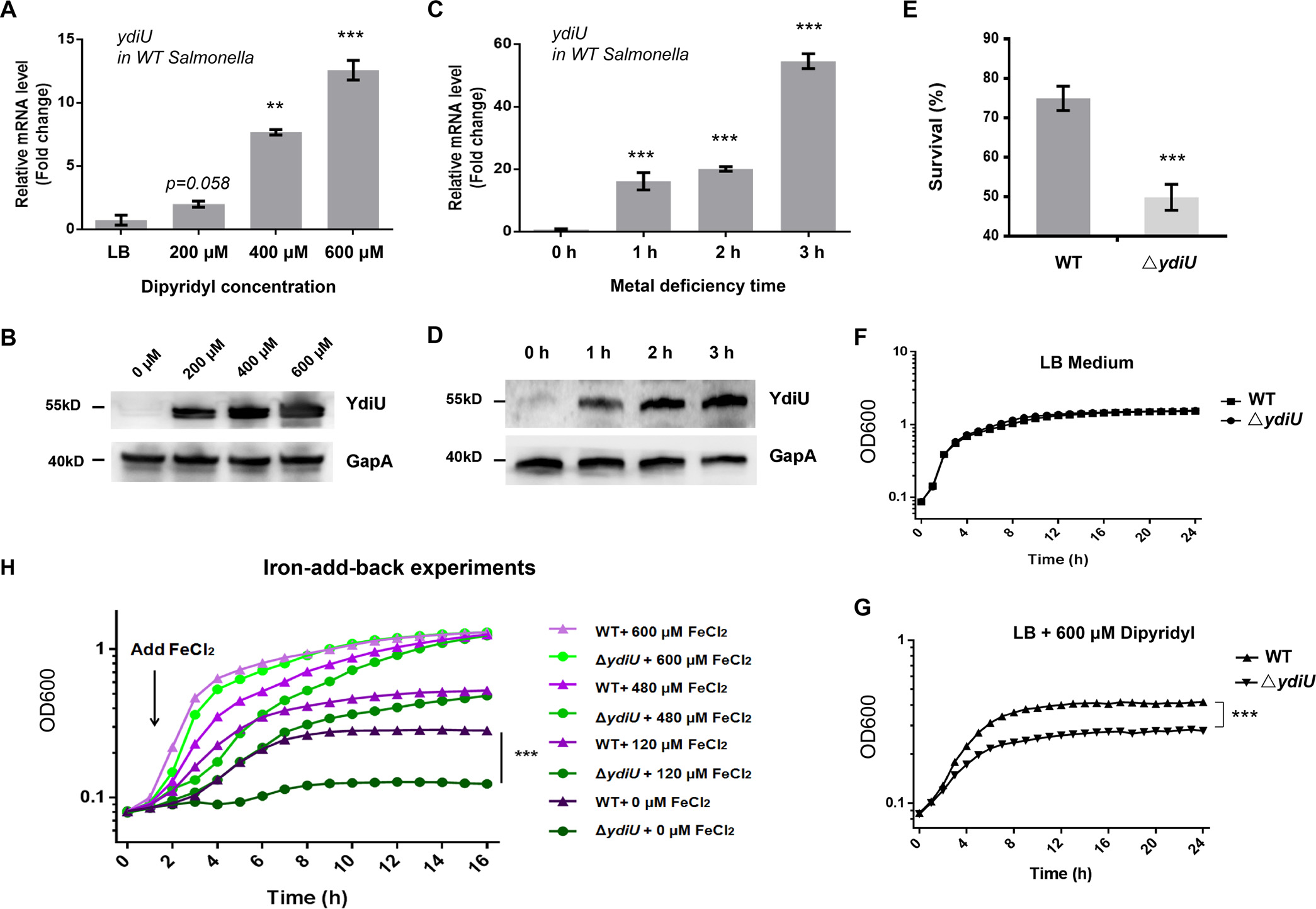
Figure 1. YdiU can be efficiently induced by metal starvation signals
Studies have shown that to combat the nutritional immunity of the host, Enterobactin (Ent) synthesized in Enterobacteriaceae is secreted out of the cell to capture iron atoms, and the Fe-bound Ents are subsequently recognized and transported into cytoplasm by a series of ferric-enterobactin uptake proteins (FepABCDEG), to improve bacterial survival in iron deficient conditions (Figure 2D). Thus the researchers, with the aid of mass spectrometry-based proteomics, found that 251 differentially expressed proteins in WT and ΔYdiU Salmonella when in metal deficient conditions (Figure 2C). Some of these Fe uptake related proteins were significantly suppressed in the ΔYdiU strain (Figure 2B). Subsequently combined with the results of qPCR (Figure 2E-F) as well as ICP-MS (Figure 2G), it was speculated that YdiU, as an activator of Fe uptake, promotes iron uptake by Salmonella in iron deficient conditions by upregulating the transcription of iron uptake relating genes, increasing the levels of proteins involved in enterobactin biosynthesis and iron-enterobactin uptake.
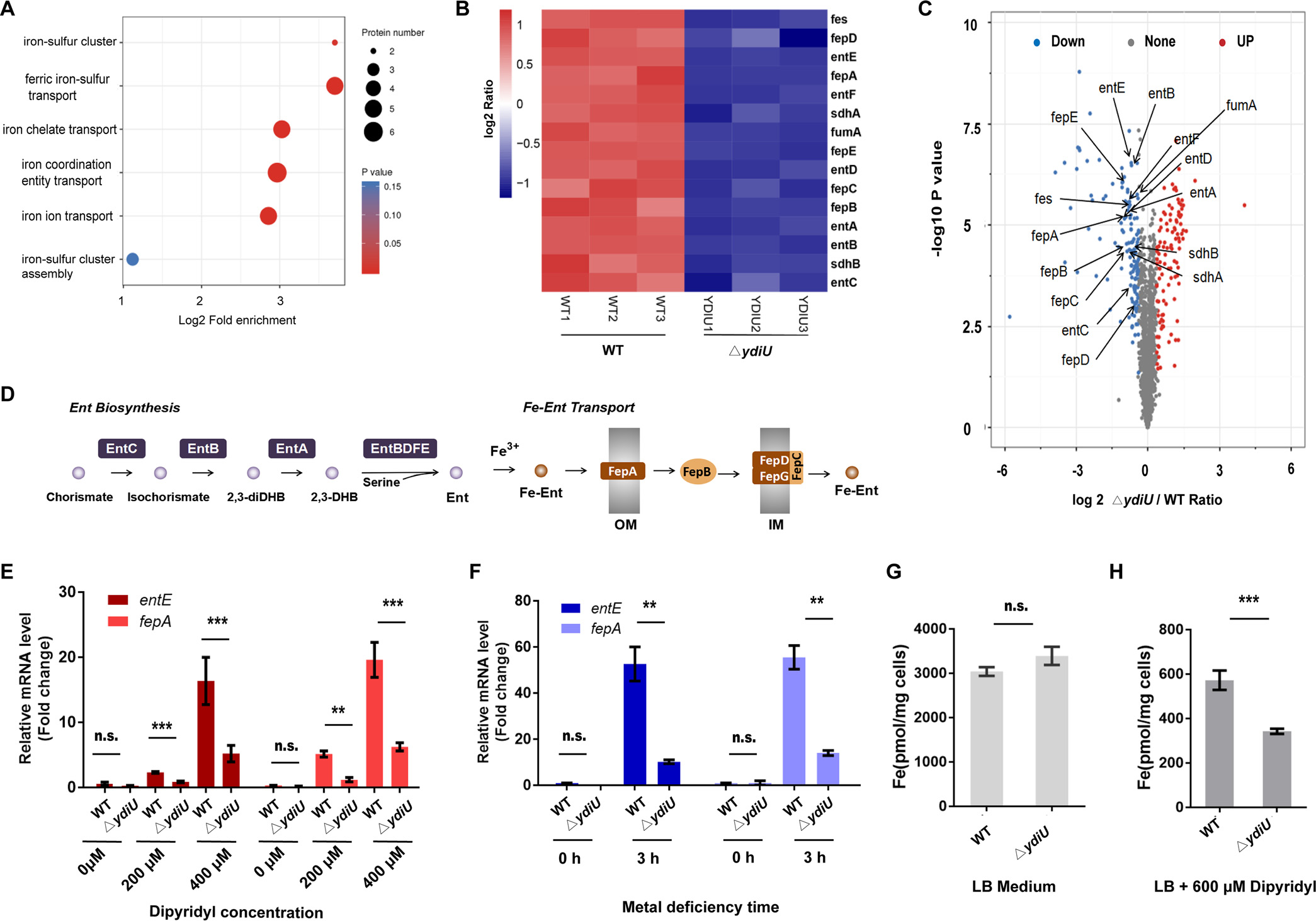
Figure 2. Proteomic and ICP-MS analysis results of WT and ΔYdiU Salmonella under metal deficient conditions
In addition to the previous study, the researchers also found in the proteomic data that the protein levels of fur, a major regulatory gene of the Salmonella iron uptake pathway, were similar between WT and ΔYdiU strains. And there was also no significant difference of their mRNA levels under equivalent conditions, determined by qPCR. This illustrates that regulation of the iron uptake pathway by YdiU must occur at a Fur post-transcriptional step. Subsequent review of the literature revealed that YdiU is an enzyme that catalyzes the post-translational modification of proteins. Previous proteomic results showed that in YdiU expressing Salmonella there are 46 UMPylated proteins, including Fur which is UMPylated at the H33 site. Since Fur is a major regulator of iron uptake, this suggests that YdiU exerts its function by directly UMPylating Fur, and additionally YdiU was found to UMPylate Fur at the H118 site.
To further investigate the function of Fur after it was UMPylated, the researchers analyzed the spatial structure of the UMPylated H33 and H118 sites, H33 locates near the Fe binding site and H118 locates on the surface of the Fur homodimer. Through sequence alignment, it was found that the UMPylated residues (H33 and H118) were highly conserved among orthologs of Fur, suggesting that the regulation of Fur by YdiU-mediated UMPylation was widely adopted. By establishing the H118A mutant (constructed by Ubigene), Fur was found to be unable to be UMPylated by YdiU, thus demonstrating that H118 is a major UMPylation site for Fur. To further investigate the effect of UMPylation on Fur characteristics, the researchers analyzed FurdYdiU (YdiU knockout) and FurpYdiU (YdiU expressed) using size exclusion chromatography, and the different elution peaks in the two samples illustrated that the aggregation state of Fur was altered after YdiU-mediated UMPylation. Subsequently, native gel assay and dynamic light scattering (DLS) were performed to compare the characteristics of native Fur and FurUMP, showing that band of FurUMP was shifted compared those of native Fur. The overall results indicate that Fur became less stable and more aggregated after UMPylation. It was subsequently verified by electrophoretic mobility shift assay (EMSA), that YdiU affected the DNA binding activity of Fur by allowing its UMPylation.
Ultimately based on the findings, the researchers proposed the following mechanistic model (Figure 3): when Salmonella are grown in an iron rich environment, the Fur protein binds to the promoter region of iron uptake genes, inhibiting the recruitment of RNA polymerase and reducing iron uptake; When Salmonella enters host cells, iron deficiency, high ROS, and low pH all activate YdiU protein expression; Then YdiU transfers one UMP group to the H118 site of Fur through UTP, preventing the dimerization of Fur; UMPylated Fur is removed from the promoters of iron uptake genes; Through this mechanism, Salmonella acquires sufficient iron to survive within host cells. The identified modification sites are highly conserved among homologous proteins, suggesting that this regulatory mechanism of Fur is widespread. The covalent attachment mode of UMP to histidine is similar to that of phosphorylated histidine (pHis), which plays a key role in prokaryotic signal transduction.
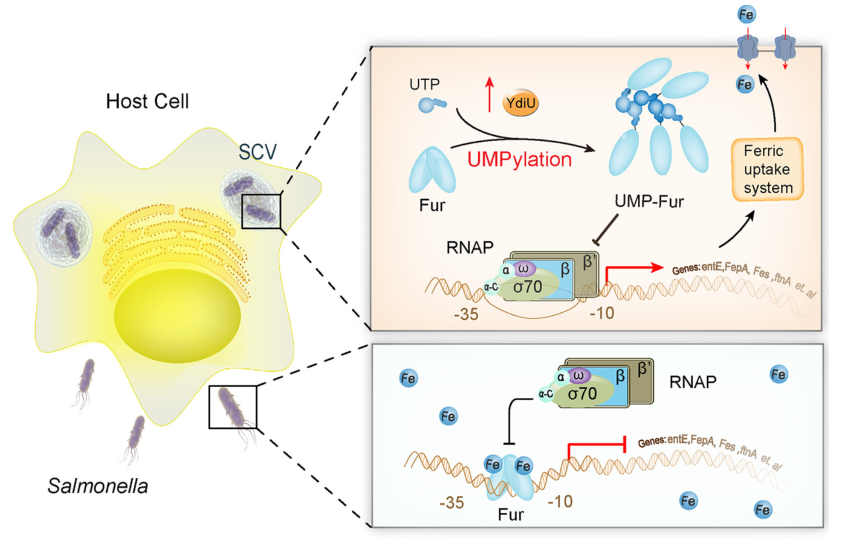
Figure 3. Model for YdiUmediated regulation of iron uptake during Salmonella infection
Ubigene took advantage of our exclusive CRISPR-B™ technology to construct the H118A mutant of Salmonella mentioned in this study which effectively verified the major UMPylation site of the iron homeostasis regulator Fur. CRISPR-B™ technology is Ubigene’s innovative technology based on Red/ET recombination system and CRISPR/Cas9 gene editing system, and by optimizing gene editing vector and gene-editing procedure, gene-editing efficiency and accuracy were increased comparing the traditional methods. This technology enables scarless gene editing of Escherichia coli/Salmonella/Pseudomonas aeruginosa and has a wide range of applications, including gene knockout, gene point mutation, and gene knock-in. Customized microbe services start from $1480, contact us for further information >>
 Subscribe Us
Subscribe Us Gene Editing Services
Gene Editing Services
 EZ-editor™
EZ-editor™ Red Cotton Gene knockout Project
Red Cotton Gene knockout Project











